Poster Session C
Pediatric autoimmune diseases: Kawasaki disease, juvenile dermatomyositis and juvenile localized scleroderma
Session: (1705–1712) Pediatric Rheumatology – Basic Science Poster
1708: In Cis SOCS1 Variants Illustrate the Precise Regulation of Interferon Signaling Needed to Prevent Autoimmunity
Tuesday, November 14, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall
- YD
Yan Du, MD, PhD
Harvard medical school
Brookline, MA, United StatesDisclosure information not submitted.
Abstract Poster Presenter(s)
Yan Du1, Evan Hsu2, Kailey Brodeur3, Meng Liu4, Mindy Lo5, Craig Platt3 and Pui Lee5, 1Harvard medical school, Boston, MA, 2Wesleyan University, Newton, MA, 3Boston Children's Hospital, Boston, MA, 4Southern Medical University, Boston, MA, 5Division of Immunology, Boston Children's Hospital, Boston, MA
Background/Purpose: Systemic autoimmunity can be driven by monogenic or polygenic risk variants. We aimed to characterize the genetic basis of disease in a family with early-onset autoimmune manifestations including systemic lupus erythematosus (SLE), discoid lupus, and immune thrombocytopenia.
Methods: Whole exome sequencing (WES) was conducted to identify candidate variants. The impact of the identified variants was analyzed by flow cytometry, immunoprecipitation, and luciferase reporter assay in transfected 293T cells. Gene expression in peripheral blood mononuclear cells was profiled by bulk RNA sequencing and plasma cytokines were measured by proximity extension assay.
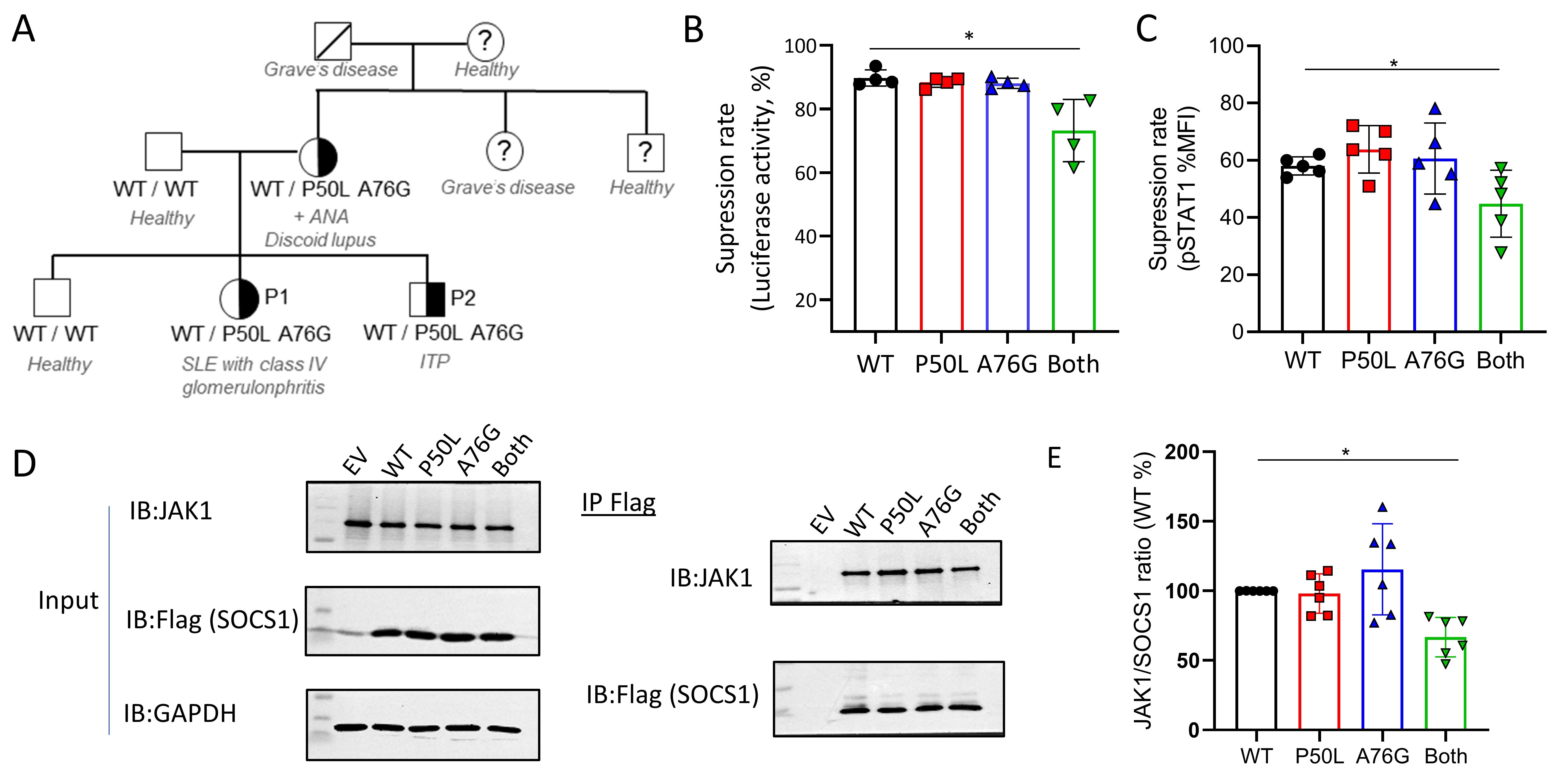
Results: We studied two siblings with early-onset systemic lupus erythematosus and immune thrombocytopenia. Mother of the probands had a history of discoid lupus and positive antinuclear antibody test. WES identified maternally inherited in cis variants (p. Pro50Leu and p.Ala76Gly) in Suppressor of cytokine signaling 1 (SOCS1) flanking the kinase inhibitory domain (Figure 1A). Both variants are predicted to be benign by in silico algorithms and neither variant alone affected the ability of SOCS1 to inhibit JAK-STAT1 signaling by luciferase reporter assay and flow cytometry quantification of STAT1 phosphorylation (Figure 1B,C). Immunoprecipitation studies showed that when both variants were expressed in cis, the mutant SOCS1 protein displayed decreased binding to JAK1 and reduced capacity to inhibit IFN-I signaling by ~25% compared to wildtype protein (Figure 1D,E).
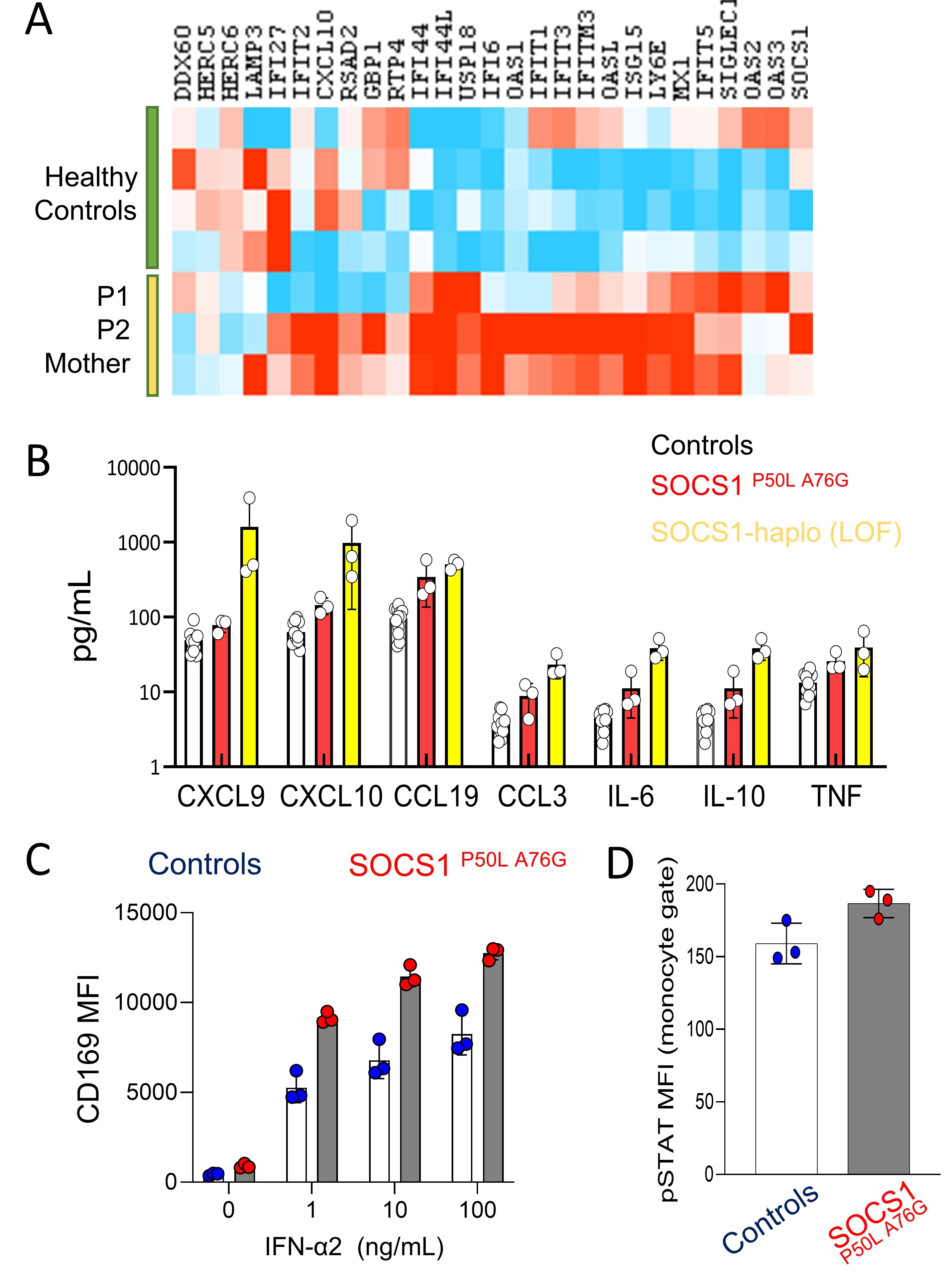
Supporting defective regulation of IFN-I, transcriptomic and cytokine analysis of both siblings and their mother showed increased expression of interferon-inducible genes compared to healthy controls, but less striking compared to patients with SOCS1 haploinsufficiency due to loss-of-function variants (Figure 2A,B). Peripheral blood monocytes from all three subjects further displayed increased CD169 expression and STAT1 phosphorylation upon IFN-I stimulation (Figure 2C,D).
Conclusion: Our work illustrates the critical fine-turning of IFN-I signaling by SOCS1 to prevent autoimmunity and demonstrates that a combination of variants that are individually benign may have deleterious consequences.
Y. Du: None; E. Hsu: None; K. Brodeur: None; M. Liu: None; M. Lo: None; C. Platt: None; P. Lee: None.
Background/Purpose: Systemic autoimmunity can be driven by monogenic or polygenic risk variants. We aimed to characterize the genetic basis of disease in a family with early-onset autoimmune manifestations including systemic lupus erythematosus (SLE), discoid lupus, and immune thrombocytopenia.
Methods: Whole exome sequencing (WES) was conducted to identify candidate variants. The impact of the identified variants was analyzed by flow cytometry, immunoprecipitation, and luciferase reporter assay in transfected 293T cells. Gene expression in peripheral blood mononuclear cells was profiled by bulk RNA sequencing and plasma cytokines were measured by proximity extension assay.
Results: We studied two siblings with early-onset systemic lupus erythematosus and immune thrombocytopenia. Mother of the probands had a history of discoid lupus and positive antinuclear antibody test. WES identified maternally inherited in cis variants (p. Pro50Leu and p.Ala76Gly) in Suppressor of cytokine signaling 1 (SOCS1) flanking the kinase inhibitory domain (Figure 1A). Both variants are predicted to be benign by in silico algorithms and neither variant alone affected the ability of SOCS1 to inhibit JAK-STAT1 signaling by luciferase reporter assay and flow cytometry quantification of STAT1 phosphorylation (Figure 1B,C). Immunoprecipitation studies showed that when both variants were expressed in cis, the mutant SOCS1 protein displayed decreased binding to JAK1 and reduced capacity to inhibit IFN-I signaling by ~25% compared to wildtype protein (Figure 1D,E).
Supporting defective regulation of IFN-I, transcriptomic and cytokine analysis of both siblings and their mother showed increased expression of interferon-inducible genes compared to healthy controls, but less striking compared to patients with SOCS1 haploinsufficiency due to loss-of-function variants (Figure 2A,B). Peripheral blood monocytes from all three subjects further displayed increased CD169 expression and STAT1 phosphorylation upon IFN-I stimulation (Figure 2C,D).
Conclusion: Our work illustrates the critical fine-turning of IFN-I signaling by SOCS1 to prevent autoimmunity and demonstrates that a combination of variants that are individually benign may have deleterious consequences.

Figure 1. A) in cis SOCS1 variants in a family with multiple autoimmune manifestations. B). Suppression of IFNα2-induced luciferase activity in ISRE-luc reporter cells by ectopic expression of wildtype and mutant SOCS1. Luciferase was measured 24 hours after IFN stimulation. C) Suppression of IFNα2-induced STAT1 phosphorylation in 293T cells by ectopic expression of wildtype and mutant SOCS1. Flow cytometry for phospho-STAT1 was performed 5 minutes after IFN stimulation. D) Assessment of SOCS1-JAK1 interaction by immunoprecipitation of Flag-tagged SOCS1 and JAK1. E) Densitometry analysis of JAK1/SOCS1 ratio from 6 independent immunoprecipitation studies. * p < 0.05.

Figure 2. A) Heatmap display of IFN-inducible genes from bulk RNAseq of PBMCs from the healthy controls, probands and their mother. B) Quantification of cytokine and chemokine levels in the plasma of healthy controls, SOCS1 mutant family and SOCS1-haplosufficiency patient by Olink proximity extension assay. C) CD169 expression on CD14+ monocytes 24 hours after stimulation with IFN-2. Three individuals with SOCS1 haploinsufficiency with confirmed loss-of-function variants were included as controls. D) Flow cytometry quantification of phospho-STAT1 in CD14+ monocytes 5 minutes after IFN-2 stimulation.
Y. Du: None; E. Hsu: None; K. Brodeur: None; M. Liu: None; M. Lo: None; C. Platt: None; P. Lee: None.



