Poster Session C
Systemic lupus erythematosus (SLE)
Session: (2257–2325) SLE – Diagnosis, Manifestations, & Outcomes Poster III
2292: Is Machine Learning Useful to Predict Flare During Pregnancy in Systemic Lupus Erythematosus?
Tuesday, November 14, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall
- EE
Elena Elefante, MD
Rheumatology Unit, Department of clinical and experimental medicine, University of Pisa
Pisa, Pisa, ItalyDisclosure information not submitted.
Abstract Poster Presenter(s)
Dina Zucchi1, Giancarlo Cascarano2, Antonio Collesei3, Elena Elefante2, Francesca Monacci4, Chiara Ietto4, Linda Carli2, Sabrina Gori2, Valentina Gelsi4, Chiara Tani2 and Marta Mosca2, 1Rheumatology Unit, Department of Clinical and Experimental Medicine, University of Pisa, Italy; Department of Medical Biotechnologies, University of Siena, Pisa, Italy, 2Rheumatology Unit, Department of Clinical and Experimental Medicine, University of Pisa, Pisa, Italy, 3Surgery, Oncology and Gastroenterology, University of Padua, Padua, Italy; Familial Cancer Clinics, Veneto Institute of Oncology, IOV-IRCCS, Padua, Italy, 4Department of Clinical and Experimental Medicine, Division of Obstetrics and Gynecology, University of Pisa, Pisa, Italy
Background/Purpose: Systemic Lupus Erythematosus (SLE) patients are at risk of disease flare during pregnancy; risk stratification is crucial for personalized treatment and monitoring. Our hypothesis is that a machine learning (ML) approach could improve the flare prediction accuracy, enabling early identification of high-risk pregnancies. In this study, we aimed to build a ML model capable of predicting disease flares based on clinical and serological data obtained early in pregnancy.
Methods: 105 pregnancies in 84 SLE patients were prospectively followed at 4-week intervals during pregnancy and puerperium. At the first visit during pregnancy, demographic and clinical data, treatments as well as disease state at the last visit before pregnancy were gathered. At each visit, disease activity was assessed by means of the Systemic Lupus Erythematosus Disease Activity Index (SLEDAI) and Physician Global Assessment (PhGA). Disease state was defined according to the LLDAS and DORIS definitions. Flares were defined as the emergence or worsening of symptoms requiring therapeutic adjustments or hospitalization. Classical inferential statistics by means of multivariate logistic regression were paralleled by a ML approach to predict the risk of flare based on baseline variables. After missing data imputation and variable filtering, the dataset was subdivided into training and test set, with 70-30% split. A Generalized Linear Model (GLM) with repeated cross-validation was then fitted on the training set, with the aim of predicting "the emergence of flares".
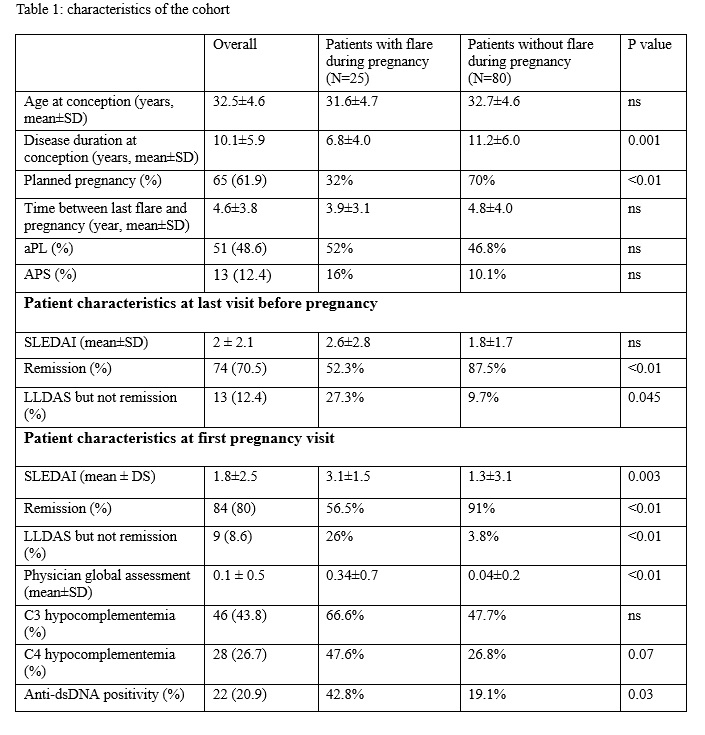
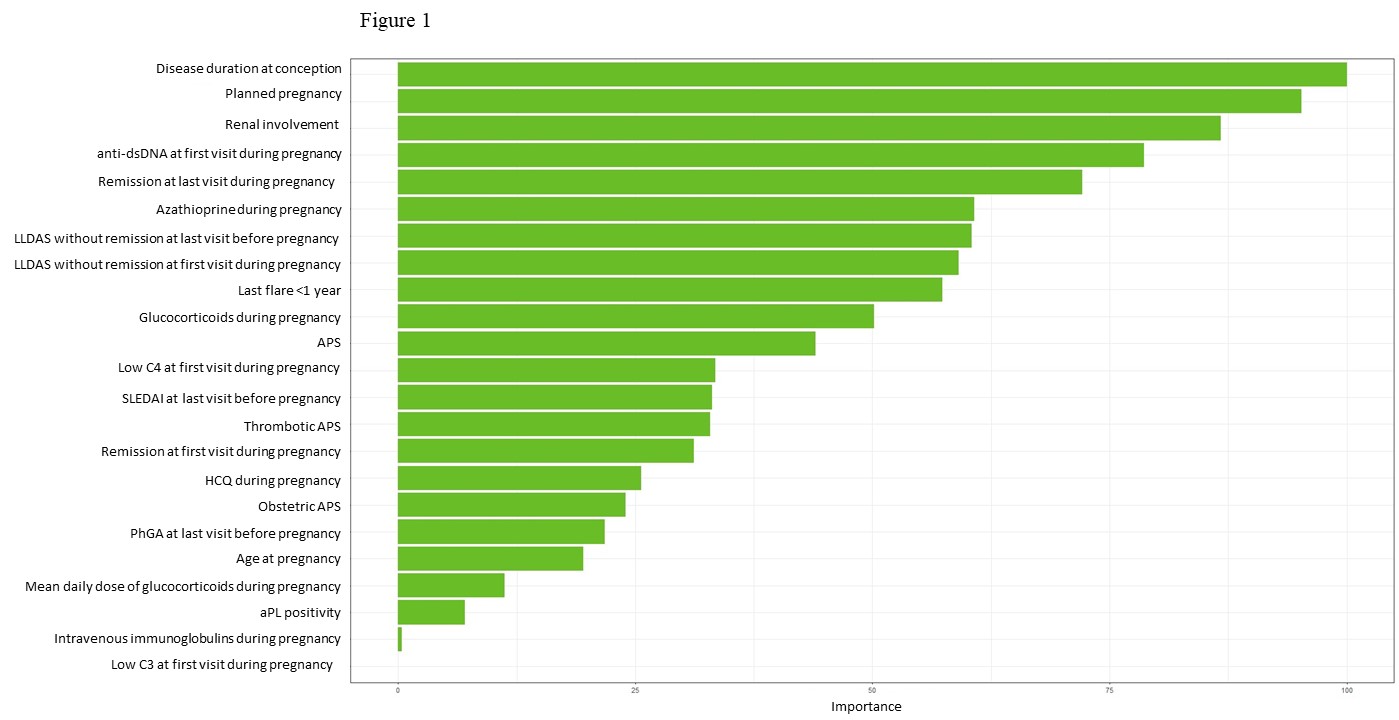
Results: 25 flares occurred among the 105 pregnancies examined (26.25%). Patients' demographic and clinical data are shown in Table 1. At univariate analysis, the variables significantly associated with flare were LLDAS without remission at last evaluation (p=0.045) and at first visit during pregnancy (p< 0.01), shorter disease duration (p< 0.01), anti-dsDNA positivity (p=0.03), higher SLEDAI (p=0.02) and higher PGA (p=0.02) at first visit during pregnancy. Remission status at both last (p< 0.01) and first visit during pregnancy (p< 0.01) and planned pregnancy (p< 0.01) were protective. In the multivariate analysis, planned pregnancy and disease duration remained significant (OR 0.84 CI 0.72 to 0.98 p=0.02 and OR 0.18 CI 0.45 to 0.78 p=0.02, respectively). The ML model achieves a high accuracy of 96.3% in predicting the target, unlocking the possibility to enact prevention measures on fragile subjects. Additionally, the model identifies key parameters for patient information collection. The model's optimization gives different weights and statistical importance to each variable (Figure 1), confirming the importance of the two previously recognized predictors, including renal involvement, among others.
Conclusion: The constructed ML model has demonstrated remarkable accuracy in identifying flare occurrences during pregnancy based on baseline data. It is worth noting that both the statistical analysis and the ML model emphasized parameters such as proper pregnancy planning and disease duration at conception as the most influential in predicting the likelihood of a flare during gestation.
D. Zucchi: None; G. Cascarano: None; A. Collesei: None; E. Elefante: None; F. Monacci: None; C. Ietto: None; L. Carli: None; S. Gori: None; V. Gelsi: None; C. Tani: None; M. Mosca: AstraZeneca, 2, Bristol-Myers Squibb(BMS), 2, Eli Lilly, 2, GlaxoSmithKlein(GSK), 2, Otsuka, 2, UCB, 2.
Background/Purpose: Systemic Lupus Erythematosus (SLE) patients are at risk of disease flare during pregnancy; risk stratification is crucial for personalized treatment and monitoring. Our hypothesis is that a machine learning (ML) approach could improve the flare prediction accuracy, enabling early identification of high-risk pregnancies. In this study, we aimed to build a ML model capable of predicting disease flares based on clinical and serological data obtained early in pregnancy.
Methods: 105 pregnancies in 84 SLE patients were prospectively followed at 4-week intervals during pregnancy and puerperium. At the first visit during pregnancy, demographic and clinical data, treatments as well as disease state at the last visit before pregnancy were gathered. At each visit, disease activity was assessed by means of the Systemic Lupus Erythematosus Disease Activity Index (SLEDAI) and Physician Global Assessment (PhGA). Disease state was defined according to the LLDAS and DORIS definitions. Flares were defined as the emergence or worsening of symptoms requiring therapeutic adjustments or hospitalization. Classical inferential statistics by means of multivariate logistic regression were paralleled by a ML approach to predict the risk of flare based on baseline variables. After missing data imputation and variable filtering, the dataset was subdivided into training and test set, with 70-30% split. A Generalized Linear Model (GLM) with repeated cross-validation was then fitted on the training set, with the aim of predicting "the emergence of flares".
Results: 25 flares occurred among the 105 pregnancies examined (26.25%). Patients' demographic and clinical data are shown in Table 1. At univariate analysis, the variables significantly associated with flare were LLDAS without remission at last evaluation (p=0.045) and at first visit during pregnancy (p< 0.01), shorter disease duration (p< 0.01), anti-dsDNA positivity (p=0.03), higher SLEDAI (p=0.02) and higher PGA (p=0.02) at first visit during pregnancy. Remission status at both last (p< 0.01) and first visit during pregnancy (p< 0.01) and planned pregnancy (p< 0.01) were protective. In the multivariate analysis, planned pregnancy and disease duration remained significant (OR 0.84 CI 0.72 to 0.98 p=0.02 and OR 0.18 CI 0.45 to 0.78 p=0.02, respectively). The ML model achieves a high accuracy of 96.3% in predicting the target, unlocking the possibility to enact prevention measures on fragile subjects. Additionally, the model identifies key parameters for patient information collection. The model's optimization gives different weights and statistical importance to each variable (Figure 1), confirming the importance of the two previously recognized predictors, including renal involvement, among others.
Conclusion: The constructed ML model has demonstrated remarkable accuracy in identifying flare occurrences during pregnancy based on baseline data. It is worth noting that both the statistical analysis and the ML model emphasized parameters such as proper pregnancy planning and disease duration at conception as the most influential in predicting the likelihood of a flare during gestation.


D. Zucchi: None; G. Cascarano: None; A. Collesei: None; E. Elefante: None; F. Monacci: None; C. Ietto: None; L. Carli: None; S. Gori: None; V. Gelsi: None; C. Tani: None; M. Mosca: AstraZeneca, 2, Bristol-Myers Squibb(BMS), 2, Eli Lilly, 2, GlaxoSmithKlein(GSK), 2, Otsuka, 2, UCB, 2.



