Poster Session C
Spondyloarthritis (SpA) including psoriatic arthritis (PsA)
Session: (1776–1795) Spondyloarthritis Including Psoriatic Arthritis – Basic Science Poster
1792: Identifying Synovial Fluid Micro-RNA Signature That Distinguishes Psoriatic Arthritis from Osteoarthritis
Tuesday, November 14, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall
- DG
Darshini Ganatra, PhD
University Health Network
Toronto, ON, CanadaDisclosure information not submitted.
Abstract Poster Presenter(s)
Darshini Ganatra1, Anas Samman1, David Nasri1, Sara Rahmati1, Omar Cruz Correa1, Rohan Machhar1, Rajiv Gandhi2, Mohit Kapoor3 and Vinod Chandran4, 1Schroeder Arthritis Institute, Krembil Research Institute, University Health Network, Toronto, ON, Canada, 2Schroeder Arthritis Institute, Krembil Research Institute, University Health Network and Division of Orthopaedics, Department of Surgery, University of Toronto, Toronto, ON, Canada, 3Schroeder Arthritis Institute, University Health Network and Department of Surgery, University of Toronto, Toronto, ON, Canada, 4Schroeder Arthritis Institute, Krembil Research Institute, University Health Network and Division of Rheumatology, Department of Medicine, University of Toronto, Toronto, ON, Canada
Background/Purpose: Psoriatic Arthritis (PsA) is an inflammatory arthritis and shares several clinical features with Osteoarthritis (OA), the most common form of arthritis. This often leads to challenges in correctly diagnosing these diseases. Synovial Fluid (SF) from joints of arthritis patients contain a rich population of pathogenic modulators, such as microRNAs (miRNAs), making it a good source to study disease biomarkers. In this study, we aimed to 1) identify differentially expressed miRNA in SF of PsA and OA patients, and 2) identify differentially expressed miRNA driven pathways.
Methods: Synovial fluid aspirated from knees of 12 PsA (sex: 8M, 4F; age (years): (median, range) = (45 (19-64)) and 12 OA patients (sex: 6M, 6F, age: (median, range) = (60 (43-74)) was used for miRNA sequencing using Illumina NextSeq 550. Linear modelling with empirical Bayes moderation (using the Limma R package) was used for assessing differential expression of miRNAs. Gene targets and enriched pathways targeted by significant miRNAs were identified by using integrative tools miRDIP and pathDIP set to a very high confidence class. For validation, SF samples from additional 35 PsA and 37 OA patients were retrieved. Total RNA including miRNA was extracted from these samples, and 5 selected candidate miRNAs were validated using quantitative RT-PCR. ΔΔCt method was used to calculate fold change. Differentially expressed miRNAs were defined as those with a two-fold change in expression (-1.0 ≥ log2FC ≥ 1.0; p-value < 0.05).
Results: After miRNA sequencing, 51 miRNAs were found to be significantly differentially expressed between PsA and OA (FDR adjusted p-value less than 0.05) (Fig. 1). QKI (RNA binding protein), OTUD4 (TLR signalling regulator), and NFAT5 (osmotic stress response regulator) in PsA, and, CELF2 (mRNA processor), NFAT5, and ACVR2B (TGFβ receptor) in OA, were the highest ranked gene targets of significant miRNAs.Figure 2 shows the signalling cascade that include genes targeted by miRNAs upregulated in PsA and OA. Pathways such as mitogen-activated protein kinase (MAPK), and WNT signalling were included which have also been previously demonstrated to be associated with PsA and OA. On qRT-PCR, 2 out of 5 miRNAs; miR-27b-3p (log2FC= -2.391, p-value < 0.05), and miR-223-3p (log2FC= 2.830, p-value < 0.05), were significantly dysregulated in SF of PsA patients compared to OA patients. Analysis using miRDIP and pathDIP revealed 195 gene targets and 114 enriched pathways for miR-27b-3p (Table 1a), and 223 gene targets and 9 enriched pathways for miR-223-3p (Table 1b).
Conclusion: Several miRNAs are deregulated between PsA and OA. miR-27b-3p, and miR-223-3p were validated in an independent cohort of patients. Identification of gene targets and enriched pathways further support the validated findings. Additional analysis of the remaining significant miRNAs will provide valuable pathogenetic insights and may identify diagnostic signature of PsA vs OA.

.jpg)
D. Ganatra: None; A. Samman: None; D. Nasri: None; S. Rahmati: None; O. Cruz Correa: None; R. Machhar: None; R. Gandhi: None; M. Kapoor: None; V. Chandran: AbbVie, 1, 5, 6, Amgen, 1, 5, 6, AstraZeneca, 3, Bristol-Myers Squibb (BMS), 1, 6, Eli Lilly, 1, 5, 6, Janssen, 1, 6, Novartis, 1, 1, 6, UCB, 1, 2.
Background/Purpose: Psoriatic Arthritis (PsA) is an inflammatory arthritis and shares several clinical features with Osteoarthritis (OA), the most common form of arthritis. This often leads to challenges in correctly diagnosing these diseases. Synovial Fluid (SF) from joints of arthritis patients contain a rich population of pathogenic modulators, such as microRNAs (miRNAs), making it a good source to study disease biomarkers. In this study, we aimed to 1) identify differentially expressed miRNA in SF of PsA and OA patients, and 2) identify differentially expressed miRNA driven pathways.
Methods: Synovial fluid aspirated from knees of 12 PsA (sex: 8M, 4F; age (years): (median, range) = (45 (19-64)) and 12 OA patients (sex: 6M, 6F, age: (median, range) = (60 (43-74)) was used for miRNA sequencing using Illumina NextSeq 550. Linear modelling with empirical Bayes moderation (using the Limma R package) was used for assessing differential expression of miRNAs. Gene targets and enriched pathways targeted by significant miRNAs were identified by using integrative tools miRDIP and pathDIP set to a very high confidence class. For validation, SF samples from additional 35 PsA and 37 OA patients were retrieved. Total RNA including miRNA was extracted from these samples, and 5 selected candidate miRNAs were validated using quantitative RT-PCR. ΔΔCt method was used to calculate fold change. Differentially expressed miRNAs were defined as those with a two-fold change in expression (-1.0 ≥ log2FC ≥ 1.0; p-value < 0.05).
Results: After miRNA sequencing, 51 miRNAs were found to be significantly differentially expressed between PsA and OA (FDR adjusted p-value less than 0.05) (Fig. 1). QKI (RNA binding protein), OTUD4 (TLR signalling regulator), and NFAT5 (osmotic stress response regulator) in PsA, and, CELF2 (mRNA processor), NFAT5, and ACVR2B (TGFβ receptor) in OA, were the highest ranked gene targets of significant miRNAs.Figure 2 shows the signalling cascade that include genes targeted by miRNAs upregulated in PsA and OA. Pathways such as mitogen-activated protein kinase (MAPK), and WNT signalling were included which have also been previously demonstrated to be associated with PsA and OA. On qRT-PCR, 2 out of 5 miRNAs; miR-27b-3p (log2FC= -2.391, p-value < 0.05), and miR-223-3p (log2FC= 2.830, p-value < 0.05), were significantly dysregulated in SF of PsA patients compared to OA patients. Analysis using miRDIP and pathDIP revealed 195 gene targets and 114 enriched pathways for miR-27b-3p (Table 1a), and 223 gene targets and 9 enriched pathways for miR-223-3p (Table 1b).
Conclusion: Several miRNAs are deregulated between PsA and OA. miR-27b-3p, and miR-223-3p were validated in an independent cohort of patients. Identification of gene targets and enriched pathways further support the validated findings. Additional analysis of the remaining significant miRNAs will provide valuable pathogenetic insights and may identify diagnostic signature of PsA vs OA.

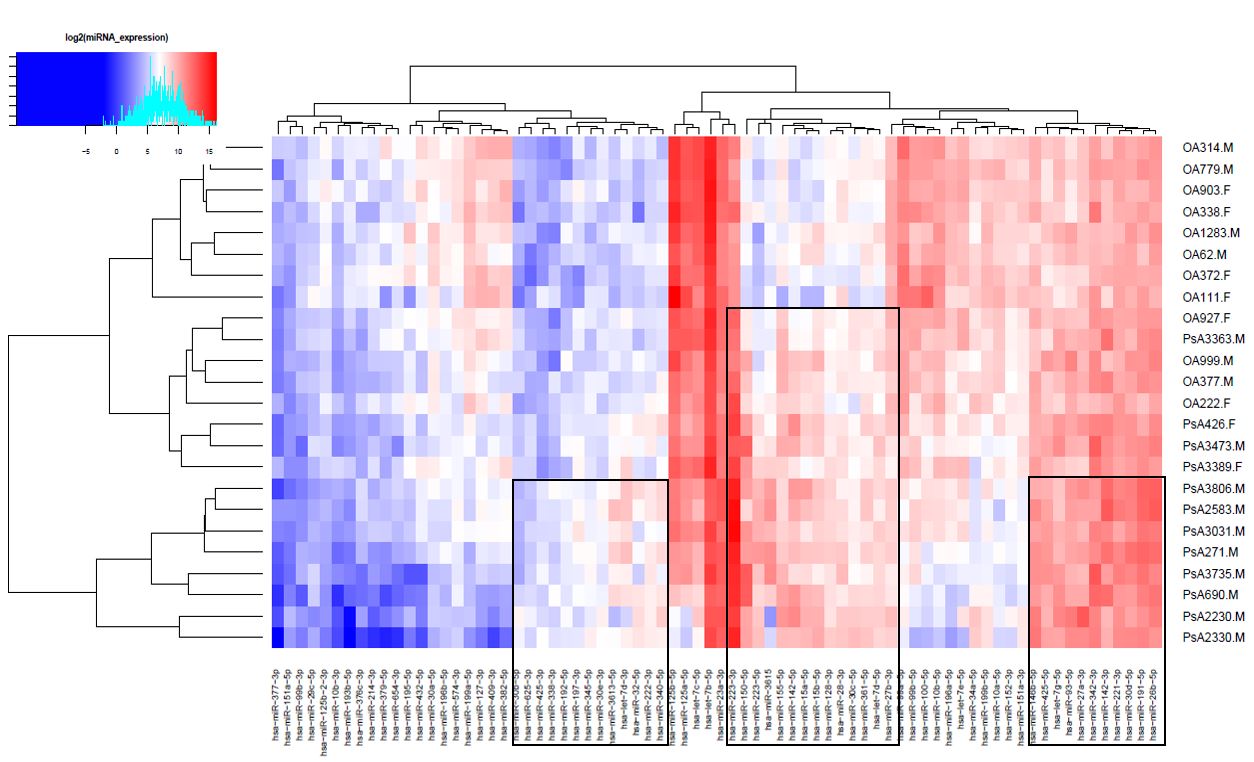
Figure. 1. Heatmap showing expression of miRNAs with Q<0.05 and fold change >1.5 in PsA and OA samples. Clusters of miRNAs that were relatively upregulated in PsA samples are indicated by the black rectangles.

Figure 2. Signaling cascades that include genes targeted by miRNAs upregulated in PsA and OA. Contextually relevant pathways were selected by applying filters for the synovium. Larger texts corresponds to highly enriched pathways.
.jpg)
Table 1. Enriched Pathways of gene targets of significant miRNAs with very high confidence of 1% and q-value <0.05. a) Enriched pathways of genes targeted by miR-27b-3p, b) Enriched pathways of genes targeted by miR-223-3p
D. Ganatra: None; A. Samman: None; D. Nasri: None; S. Rahmati: None; O. Cruz Correa: None; R. Machhar: None; R. Gandhi: None; M. Kapoor: None; V. Chandran: AbbVie, 1, 5, 6, Amgen, 1, 5, 6, AstraZeneca, 3, Bristol-Myers Squibb (BMS), 1, 6, Eli Lilly, 1, 5, 6, Janssen, 1, 6, Novartis, 1, 1, 6, UCB, 1, 2.



