Poster Session B
Pediatric autoimmune diseases: Kawasaki disease, juvenile dermatomyositis and juvenile localized scleroderma
Session: (1221–1255) Pediatric Rheumatology – Clinical Poster II: Connective Tissue Disease
1221: Familial Clustering of Dysbiotic Oral and Fecal Microbiomes in Juvenile Dermatomyositis (JDM)
Monday, November 13, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall
- AC
Albert Chow, MD (he/him/his)
Loma Linda University
Loma Linda, CA, United StatesDisclosure information not submitted.
Abstract Poster Presenter(s)
Albert Chow1, Sean Koester2, Evan Pepper-Tunick3, Peggy Lee4, Mary Eckert5, Laurie Brenchley6, Pamela Gardner6, Naisi Li2, adam Schiffenbauer7, Rita Volochayev7, Nastaran Bayat8, Jeffrey McLean9, Lisa Rider10, Susan Shenoi11, Anne Stevens12 and Neelendu Dey13, 1Loma Linda University, Loma Linda, CA, 2Translational Science and Therapeutics Division, Fred Hutchinson Cancer Center, Seattle, WA, 3Molecular Engineering & Sciences Institute, University of Washington, Seattle, WA, 4Department of Oral Medicine, School of Dentistry, University of Washington, Seattle, WA, 5Center for Clinical and Translational Research, Seattle Children’s Research Institute, Seattle, WA, 6Office of the Clinical Director, NIDCR, National Institutes of Health, Bethesda, MD, 7Environmental Autoimmunity Group, Clinical Research Branch, National Institute of Environmental Health Sciences, National Institutes of Health, Bethesda, MD, 8Social Scientific Systems, DLH Holdings Corp, Silver Spring, MD, 9Department of Periodontics, University of Washington, Seattle, WA, 10NIEHS, NIH, Bethesda, MD, 11Seattle Childrens Hospital, Mercer Island, WA, 12Janssen, Hansville, WA, 13Department of Medicine, Division of Gastroenterology, University of Washington, Seattle, WA
Background/Purpose: JDM is a rare immune-mediated disease of childhood that is thought to result from genetic predisposition and environmental drivers, with documented links to microbial exposures. In this multi-center, prospective, observational cohort study, we evaluated whether JDM is associated with discrete oral and gut microbiome signatures using siblings to control for family-associated microbial communities.
Methods: We generated 16S rRNA sequence data from fecal, saliva, supragingival, and subgingival plaque samples from JDM probands (n=28, age range 3-18 years, mean age 10 years, 46% female). To control for genetic and environmental determinants of microbiome community structure, we also profiled microbiomes of their unaffected family members (n=27 siblings, n=26 mothers, and n=17 fathers). We performed paired within-family comparisons as well as unpaired analyses of different cohorts.
Results: Family unit was the predominant factor explaining variance in both weighted and unweighted UniFrac distances between samples for each sample type (p< 0.001 in fecal, saliva, supragingival swab, and posterior subgingival dental plaque samples; p< 0.01 in anterior subgingival dental plaque samples, permutational multivariate analysis of variance. Unweighted UniFrac distances between siblings was significantly smaller than the distances between JDM probands in our cohort (p< 0.002, two-tailed Student's t-test).
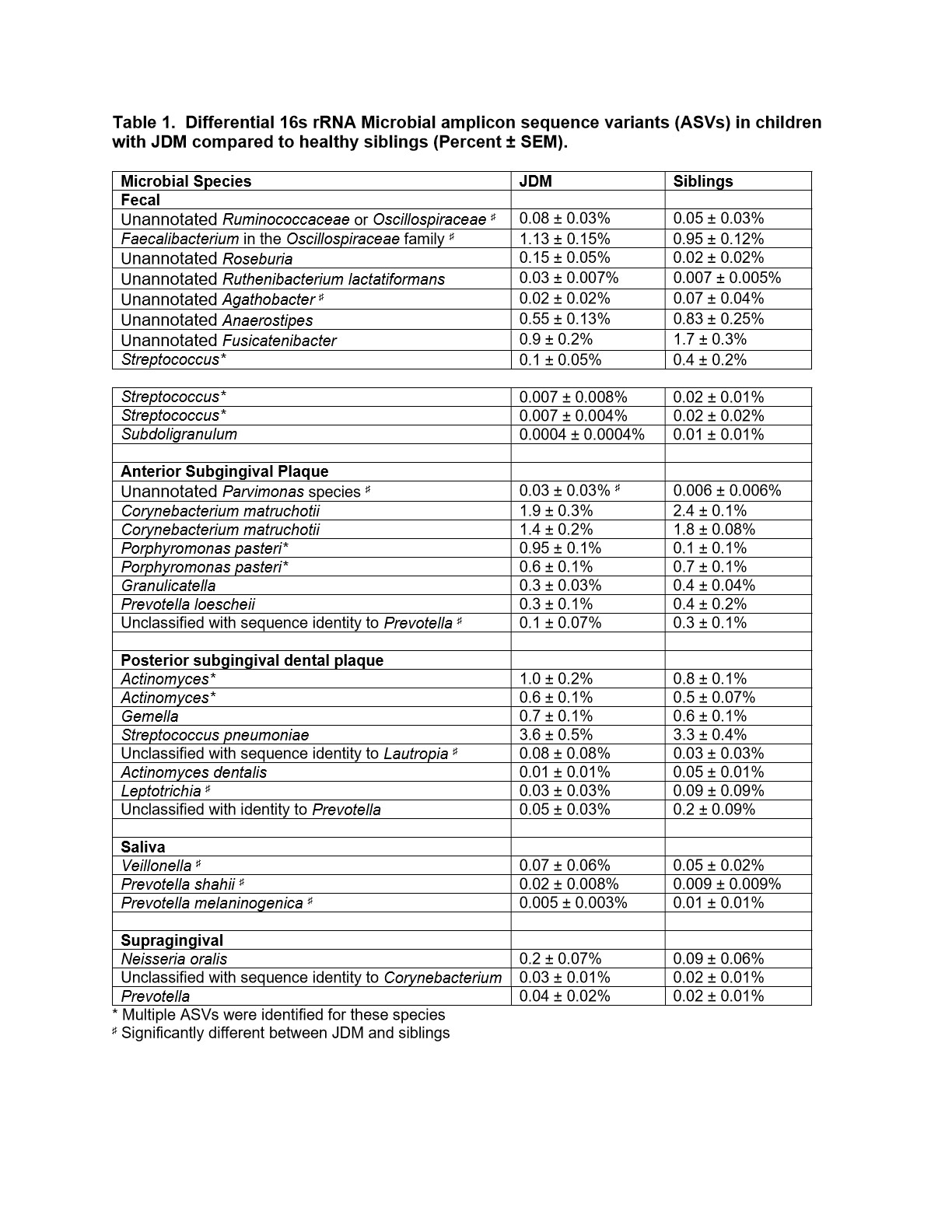
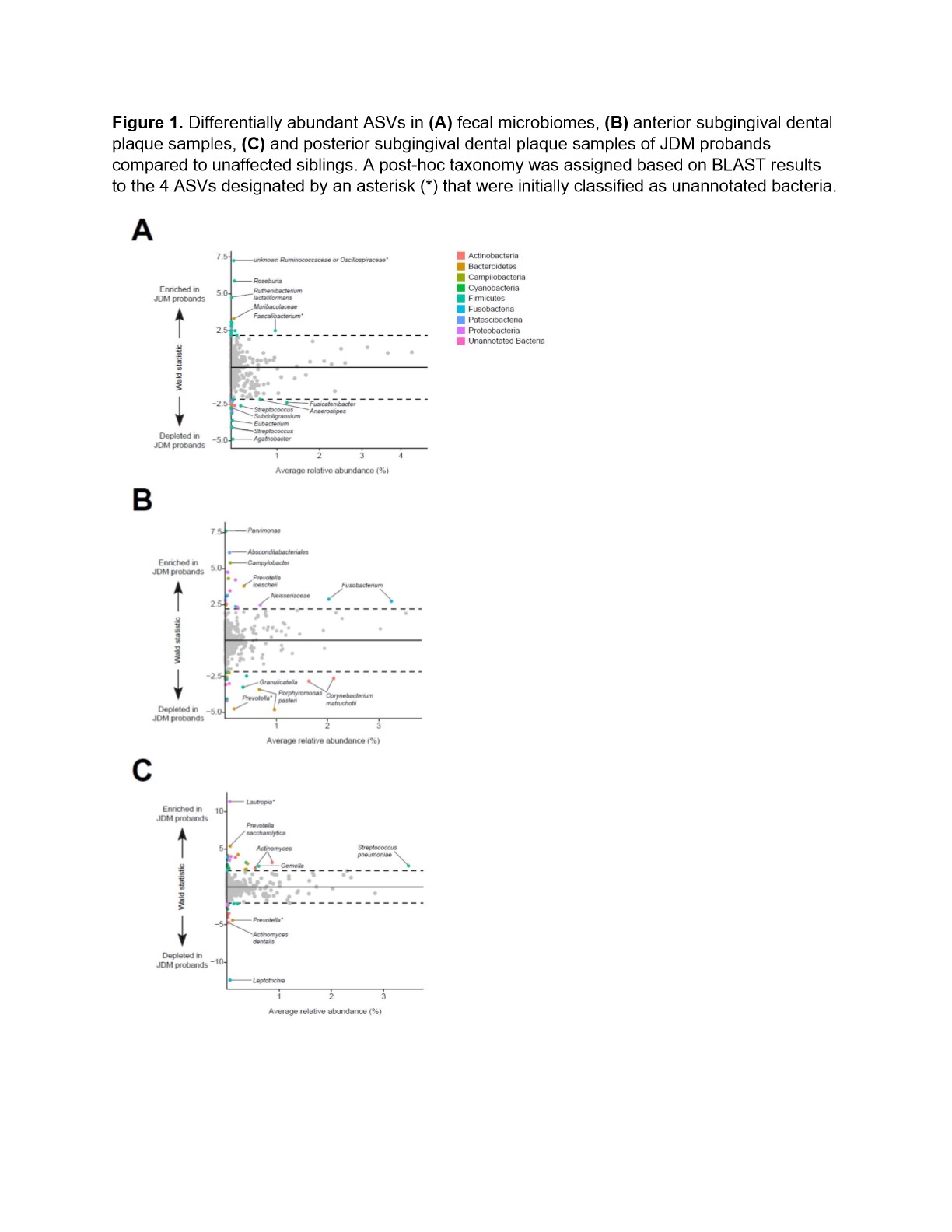
In fecal samples, 4% of bacterial amplicon sequence variants (ASVs) were differentially abundant in microbiomes between JDM probands and their siblings (Figure 1A, Table 1). In the case where ASVs that were initially classified as unannotated bacteria during the analyses, taxonomy was assigned based on BLAST results.Approximately 8% of ASVs identified in the anterior subgingival dental plaque were significantly different between JDM probands and their healthy siblings (Figure 1B). In posterior subgingival dental plaque samples, approximately 7% of identified ASVs were significantly differentially abundant between JDM probands and their siblings (Figure 1C). We also observed differentially abundant ASVs in the saliva and supragingival swab samples from JDM probands, all of which were found in very low abundances, including salivary enrichment of Veillonella and Prevotella shahii and depletion of Prevotellamelaninogenica.
Conclusion: The oral and gut microbiomes of JDM probands were more similar to their own unaffected siblings than they were to the microbiomes of other JDM probands, suggesting family unit has a significant effect on microbiome community structure. Nonetheless, in a sibling-paired analysis, several potentially immunomodulatory bacterial taxa were differentially abundant in the microbiomes of JDM probands, including Faecalibacterium in the gut and Streptococcus in the oral cavity. The loss or gain of specific fecal and oral bacteria may potentially play a role in disease pathogenesis and/or be secondary to immune dysfunction in susceptible individuals.
A. Chow: None; S. Koester: None; E. Pepper-Tunick: None; P. Lee: None; M. Eckert: None; L. Brenchley: None; P. Gardner: None; N. Li: None; a. Schiffenbauer: None; R. Volochayev: None; N. Bayat: None; J. McLean: None; L. Rider: AstraZeneca, 5, Bristol-Myers Squibb(BMS), 5, Hope Pharmaceuticals, 5; S. Shenoi: None; A. Stevens: Janssen, 2, 5; N. Dey: None.
Background/Purpose: JDM is a rare immune-mediated disease of childhood that is thought to result from genetic predisposition and environmental drivers, with documented links to microbial exposures. In this multi-center, prospective, observational cohort study, we evaluated whether JDM is associated with discrete oral and gut microbiome signatures using siblings to control for family-associated microbial communities.
Methods: We generated 16S rRNA sequence data from fecal, saliva, supragingival, and subgingival plaque samples from JDM probands (n=28, age range 3-18 years, mean age 10 years, 46% female). To control for genetic and environmental determinants of microbiome community structure, we also profiled microbiomes of their unaffected family members (n=27 siblings, n=26 mothers, and n=17 fathers). We performed paired within-family comparisons as well as unpaired analyses of different cohorts.
Results: Family unit was the predominant factor explaining variance in both weighted and unweighted UniFrac distances between samples for each sample type (p< 0.001 in fecal, saliva, supragingival swab, and posterior subgingival dental plaque samples; p< 0.01 in anterior subgingival dental plaque samples, permutational multivariate analysis of variance. Unweighted UniFrac distances between siblings was significantly smaller than the distances between JDM probands in our cohort (p< 0.002, two-tailed Student's t-test).
In fecal samples, 4% of bacterial amplicon sequence variants (ASVs) were differentially abundant in microbiomes between JDM probands and their siblings (Figure 1A, Table 1). In the case where ASVs that were initially classified as unannotated bacteria during the analyses, taxonomy was assigned based on BLAST results.Approximately 8% of ASVs identified in the anterior subgingival dental plaque were significantly different between JDM probands and their healthy siblings (Figure 1B). In posterior subgingival dental plaque samples, approximately 7% of identified ASVs were significantly differentially abundant between JDM probands and their siblings (Figure 1C). We also observed differentially abundant ASVs in the saliva and supragingival swab samples from JDM probands, all of which were found in very low abundances, including salivary enrichment of Veillonella and Prevotella shahii and depletion of Prevotellamelaninogenica.
Conclusion: The oral and gut microbiomes of JDM probands were more similar to their own unaffected siblings than they were to the microbiomes of other JDM probands, suggesting family unit has a significant effect on microbiome community structure. Nonetheless, in a sibling-paired analysis, several potentially immunomodulatory bacterial taxa were differentially abundant in the microbiomes of JDM probands, including Faecalibacterium in the gut and Streptococcus in the oral cavity. The loss or gain of specific fecal and oral bacteria may potentially play a role in disease pathogenesis and/or be secondary to immune dysfunction in susceptible individuals.

Table 1. Differential 16s rRNA Microbial amplicon sequence variants (ASVs) in children with JDM compared to healthy siblings (Percent ± SEM).

Figure 1. Differentially abundant ASVs in (A) fecal microbiomes, (B) anterior subgingival dental plaque samples, (C) and posterior subgingival dental plaque samples of JDM probands compared to unaffected siblings. A post-hoc taxonomy was assigned based on BLAST results to the 4 ASVs designated by an asterisk (*) that were initially classified as unannotated bacteria.
A. Chow: None; S. Koester: None; E. Pepper-Tunick: None; P. Lee: None; M. Eckert: None; L. Brenchley: None; P. Gardner: None; N. Li: None; a. Schiffenbauer: None; R. Volochayev: None; N. Bayat: None; J. McLean: None; L. Rider: AstraZeneca, 5, Bristol-Myers Squibb(BMS), 5, Hope Pharmaceuticals, 5; S. Shenoi: None; A. Stevens: Janssen, 2, 5; N. Dey: None.



