Poster Session B
Fibrosing rheumatic diseases (scleroderma, MCTD, IgG4-related disease, scleroderma mimics)
Session: (0934–0964) Systemic Sclerosis & Related Disorders – Basic Science Poster
0941: RUNX1 Expression and Binding Site Accessibility Is Associated with Increased Disease Severity in Systemic Sclerosis
Monday, November 13, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall
- RP
Rezvan Parvizi, PharmD, MSc
Dartmouth
lebanon, NH, United StatesDisclosure information not submitted.
Abstract Poster Presenter(s)
Rezvan Parvizi1, Zhiyun Gong1, Dillon Popovich1, Tamar Abel2, Helen Jarnagin1, Madeline Morrisson1, Tammara Wood2, Jonathan Garlick3, Monique Hinchcliff4, Patricia Pioli1 and Michael Whitfield1, 1Geisel School of Medicine at Dartmouth, Hanover, NH, 2Dartmouth College, Hanover, NH, 3Tufts University School of Dental Medicine, Boston, MA, 4Yale School of Medicine, Westport, CT
Background/Purpose: Runt-related transcription factor 1 (RUNX1) is a member of the core-binding factor family that regulates proliferation, differentiation, and cell survival in multiple cell lineages. Activation of RUNX1 in fibroblasts has been implicated in wound healing and a network analysis of transcription factor (TF) activity suggested that it is a key regulator in diffuse cutaneous systemic sclerosis (dcSSc) skin. Here, we analyze the specific contribution and function of RUNX1 in systemic sclerosis (SSc) dermal fibroblasts.
Methods:
Results: Analysis of gene expression data from a cohort of 124 individuals revealed significant over-expression of RUNX1 in skin biopsies of SSc patients. Correlation analysis demonstrated significantly increased expression of RUNX1 and in patients with higher modified Rodnan Skin Score (mRSS) (r2= 0.33, p-value = 4.4e-10) (Fig 1A, B). Analysis of scRNA-seq data from SSc and control skin showed enrichment of RUNX1 in PDGFRA1/COL1A1 expressing fibroblasts subpopulations (Fig 1C). Analysis of scRNA-seq and scATAC-seq of in vitro 3D skin-like tissues showed increase expression and RUNX1 binding site accessibility in ECM producing fibroblasts. DNA methylation profiling of isolated fibroblasts from dcSSc and healthy donors demonstrated hypomethylation of RUNX1 in SSc fibroblasts (Fig 1D). Bulk ATAC-seq of dcSSc fibroblasts showed increased accessibility of RUNX1 chromatin binding (Fig 1E), which facilitates TF binding. Inhibition of RUNX1 activity by Ro5-3335 significantly reduced the ability of dermal fibroblasts to contract collagen gel matrices (Fig 1F) and reduced fibroblast proliferation rate (Fig 1G). RUNX1 knockdown by siRNA resulted in statistically significant changes in αSMA, fibronectin (FN1) and COL1A1 expression.
Conclusion: These findings implicate RUNX1 as a key regulator of ECM producing SSc fibroblasts. Our observations indicate a potential association between the severity of dermal fibrosis and RUNX1 expression levels. Identifying hypomethylated CpG sites near the RUNX1 gene suggests their potential involvement in the upregulation of RUNX1 expression. Notably, our analysis of ATAC-seq data revealed higher accessibility of the chromatin region where RUNX1 binding sites are located. Lastly, inhibition of RUNX1 activity using Ro5-3335 caused a reduction in fibroblasts’ contraction and proliferation rates, and the regulation of key genes implicated in SSc pathogenesis.
R. Parvizi: None; Z. Gong: None; D. Popovich: None; T. Abel: None; H. Jarnagin: None; M. Morrisson: None; T. Wood: None; J. Garlick: None; M. Hinchcliff: Boehringer-Ingelheim, 5, Kadmon, 5; P. Pioli: Boehringer-Ingelheim, 1, Bristol-Myers Squibb(BMS), 1, 2, 5, Celdara Medical LLC, 2, 5, Pfizer, 5; M. Whitfield: Boehringer-Ingelheim, 1, Bristol-Myers Squibb(BMS), 2, 5, Celdara Medical, 2, 5, 12, Scientific Founder.
Background/Purpose: Runt-related transcription factor 1 (RUNX1) is a member of the core-binding factor family that regulates proliferation, differentiation, and cell survival in multiple cell lineages. Activation of RUNX1 in fibroblasts has been implicated in wound healing and a network analysis of transcription factor (TF) activity suggested that it is a key regulator in diffuse cutaneous systemic sclerosis (dcSSc) skin. Here, we analyze the specific contribution and function of RUNX1 in systemic sclerosis (SSc) dermal fibroblasts.
Methods:
RUNX1 was analyzed in gene expression data from SSc forearm skin biopsies (GSE59787) and in single-cell RNA sequencing (scRNA-seq) data from Tabib et al. 2021 (GSE138669), and in SSc and control scRNA-seq and scATAC-seq data of 3D skin-like tissues. We further generated and analyzed genome-wide DNA methylation profiles and bulk ATAC-seq data to evaluate epigenetic state and genome chromatin accessibility of RUNX1 in dcSSc-isolated fibroblasts. Fibroblasts in 2D and 3D skin-like tissues were treated with a selective RUNX1 inhibitor, Ro5-3335 or siRNA against RUNX1 followed by collagen contraction, proliferation assays and qRT-PCR for select target genes.
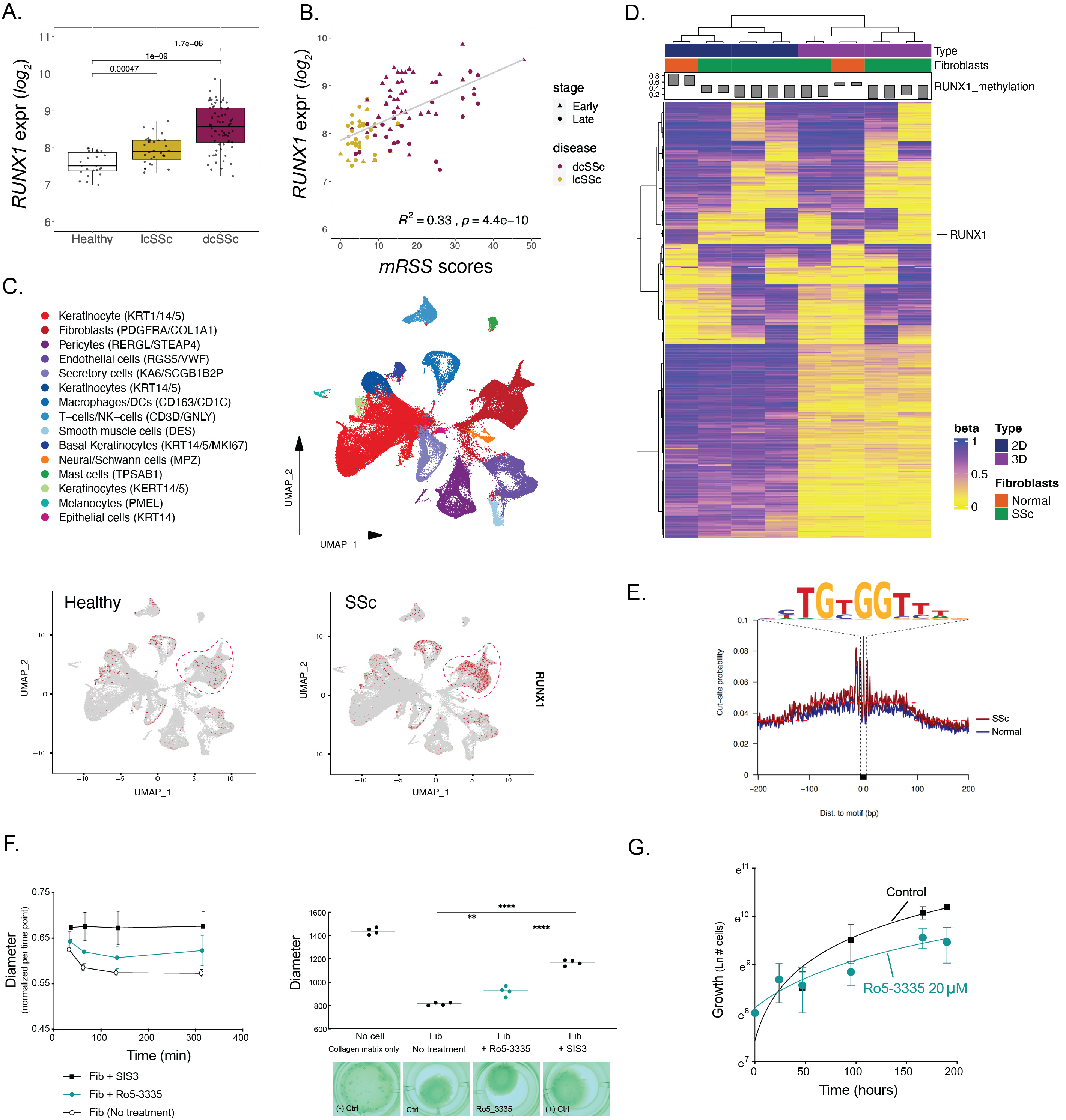
Results: Analysis of gene expression data from a cohort of 124 individuals revealed significant over-expression of RUNX1 in skin biopsies of SSc patients. Correlation analysis demonstrated significantly increased expression of RUNX1 and in patients with higher modified Rodnan Skin Score (mRSS) (r2= 0.33, p-value = 4.4e-10) (Fig 1A, B). Analysis of scRNA-seq data from SSc and control skin showed enrichment of RUNX1 in PDGFRA1/COL1A1 expressing fibroblasts subpopulations (Fig 1C). Analysis of scRNA-seq and scATAC-seq of in vitro 3D skin-like tissues showed increase expression and RUNX1 binding site accessibility in ECM producing fibroblasts. DNA methylation profiling of isolated fibroblasts from dcSSc and healthy donors demonstrated hypomethylation of RUNX1 in SSc fibroblasts (Fig 1D). Bulk ATAC-seq of dcSSc fibroblasts showed increased accessibility of RUNX1 chromatin binding (Fig 1E), which facilitates TF binding. Inhibition of RUNX1 activity by Ro5-3335 significantly reduced the ability of dermal fibroblasts to contract collagen gel matrices (Fig 1F) and reduced fibroblast proliferation rate (Fig 1G). RUNX1 knockdown by siRNA resulted in statistically significant changes in αSMA, fibronectin (FN1) and COL1A1 expression.
Conclusion: These findings implicate RUNX1 as a key regulator of ECM producing SSc fibroblasts. Our observations indicate a potential association between the severity of dermal fibrosis and RUNX1 expression levels. Identifying hypomethylated CpG sites near the RUNX1 gene suggests their potential involvement in the upregulation of RUNX1 expression. Notably, our analysis of ATAC-seq data revealed higher accessibility of the chromatin region where RUNX1 binding sites are located. Lastly, inhibition of RUNX1 activity using Ro5-3335 caused a reduction in fibroblasts’ contraction and proliferation rates, and the regulation of key genes implicated in SSc pathogenesis.

Figure 1. RUNX1 in Systemic Sclerosis Dermal Fibroblasts. A) RUNX1 expression in forearm skin biopsies of 24 healthy, 68 diffuse cutaneous systemic sclerosis (dcSSc), and 32 limited cutaneous systemic sclerosis (lcSSc) individuals. B) Pearson Correlation between RUNX1 expression and mRSS. Stage: Early, < 2 years; Late, >2 years of disease onset/diagnosis. C) UMAP projection of cell types from Tabib et al. 2021 single-cell RNA-Seq data from 10 healthy and 12 dcSSc forearm skin biopsies. Feature plots of RUNX1 is shown in heathy (left) and SSc (right) in red. Dash lines indicate fibroblast cluster. D) DNA methylation profile of 2D and 3D cultured fibroblasts that were isolated from dcSSc and healthy donors using Illumina’s Infinium Methylation EPIC array. Heatmap shows top 593 methylated CpG sits with blue/yellow gradient of beta values. The bar-plot on top shows RUNX1 beta value that is labeled within the heatmap. E) Footprint profiles of RUNX1 in SSc (red) and healthy (blue) isolated fibroblasts. The consensus sequence of RUNX1 motif is shown. F) 3D collagen contraction assays, fixed (left) and floating (right) models, of normal human dermal fibroblasts treated with Ro5-3335 (RUNX1 inhibitor). SIS3 (SMAD3 inhibitor) was used as positive control and significantly eliminates the contraction ability of fibroblasts. (Student t-test P value: ** 0.001 to 0.01, **** <0.0001 in GraphPad Prism v.9). G) Normal human dermal fibroblast proliferation curve in presence and absence of Ro5-3335.
R. Parvizi: None; Z. Gong: None; D. Popovich: None; T. Abel: None; H. Jarnagin: None; M. Morrisson: None; T. Wood: None; J. Garlick: None; M. Hinchcliff: Boehringer-Ingelheim, 5, Kadmon, 5; P. Pioli: Boehringer-Ingelheim, 1, Bristol-Myers Squibb(BMS), 1, 2, 5, Celdara Medical LLC, 2, 5, Pfizer, 5; M. Whitfield: Boehringer-Ingelheim, 1, Bristol-Myers Squibb(BMS), 2, 5, Celdara Medical, 2, 5, 12, Scientific Founder.



