Poster Session A
Rheumatoid arthritis (RA)
Session: (0423–0459) RA – Treatment Poster I
0449: Parsing Pathophysiology of Rheumatoid Arthritis-associated Lymphoproliferative Disorders via Whole RNA Transcriptome Analysis: Multi-center Study in Japan
Sunday, November 12, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall
- AT
Atsuko Tsujii, MD
Osaka Minami Medical Center
Kawachinagano City, JapanDisclosure information not submitted.
Abstract Poster Presenter(s)
ATSUKO TSUJII1, KAZUKO SAKAI2, SHIRO OHSHIMA1, YUKIHIKO SAEKI1, MASATO YAGITA3, TOMOYA MIYAMURA4, Masao Katayama5, YASUSHI HIRAMATSU6, SHINJI HIGA7, FUMINORI HIRANO8, KENJI ICHIKAWA9, NORIYUKI CHIBA10, TAKAO SUGIYAMA11, ATSUSHI IHATA12, HIROSHI TSUTANI13, KOICHIRO TAKAHI14, KIYOSHI MIGITA15, SHUNSUKE MORI16, NORIE YOSHIKAWA17, ATSUHISA UEDA18, SHOUHEI NAGAOKA19, KEIGO SETOGUCHI20, SHOJI SUGII21, ASAMI ABE22, TOSHIAKI SUGAYA23, HIROYUKI SUGAHARA24, SHINICHIRO TSUNODA24, NORISHIGE IIZUKA25, RYOSUKE YOSHIHARA26, HIROKI YABE27, TOMOAKI FUJISAKI28, EIICHI MORII29, KAZUYOSHI SAITO30, Kiyoshi Matsui31, YASUHIKO TOMITA32, HIROSHI FURUKAWA33, Shigeto Tohma34, KAZUTO NISHIO2 and YOSHIHIKO HOSHIDA35, 1National Hospital Organization (NHO), Osaka Minami Medical Center, Kawachinagano, Japan, 2Kindai University School of Medicine Department of Genome Biology, Sayama, Japan, 3Medical Research Institute KITANO HOSPITAL, PIIF Tazuke-kofukai, Osaka, Japan, 4NHO Kyushu Medical Center, Fukuoka, Japan, 5National Hospital Organization, Nagoya Medical Center, Nagoya, JP, Nagoya, Japan, 6Japanese Red Cross Society Himeji Hospital, Himeji, Japan, 7Daini Osaka Police Hospital, Osaka, Japan, 8NHO Asahikawa Medical Center, Asahikawa, Japan, 9Nissei hospital, Sapporo, Japan, 10NHO Morioka Medical Center, Morioka, Japan, 11NHO Shimoshizu Hospital, Yotsukaido, Japan, 12NHO Yokohama Medical Center, Yokohama, Japan, 13NHO Awara Hospital, Awara, Japan, 14NHO Osaka Toneyama Medical Center, Toyonaka, Japan, 15Department of Rheumatology Fukushima Medical University School of Medicine, Fukushima, Japan, 16NHO Kumamoto Saishun Medical Center, Koshi, Japan, 17NHO Miyakonojo Medical Center, Miyakonojo, Japan, 18Yokohama City University Medical Center, Yokohama, Japan, 19Yokohama Minami Kyosai Hospital, Yokohama, Japan, 20Tokyo Metropolitan Cancer and Infectious diseases Center Komagome Hospital / Tokyo Metropolitan Komagome Hospital, Tokyo, Japan, 21Tokyo Metropolitan Matsuzawa Hospital, Tokyo, Japan, 22Niigata Rheumatic Center, Shibata, Japan, 23Fuchu Hospital, Izumi, Japan, 24Sumitomo Hospital, Osaka, Japan, 25Kishiwada City Hospital, Kishiwada, Japan, 26Hyogo Prefectural Kakogawa Hospital, Kakogawa, Japan, 27Ako Central Hospital, Ako, Japan, 28Matsuyama Red Cross Hospital, Matsuyama, Japan, 29Osaka University, Suita, Japan, 30University of Occupational and Environmental Health, Kitakyushu, Japan, 31Hyogo Medical University, Nishinomiya, Japan, 32International University of Health and Welfare, Otawara City, Japan, 33NHO Tokyo National Hospital, Kiyose, Japan, 34NHO Tokyo National Hospital, Dallas, TX, 35National Hospital Organization Osaka Minami Medical Center, Kawachinagano, Japan
Background/Purpose: Lymphoproliferative disorders (LPD) in rheumatoid arthritis (RA) (RA-LPD) have unique pathophysiological features. More than half of the cases of RA-LPD undergo spontaneous regression after the discontinuation of disease modifying anti rheumatic drugs (DMARDs), whereas approximately 33% of these cases relapse. Epstein-Barr virus (EBV) infection is especially involved in the pathophysiology of RA-LPD. The current study was conducted with whole RNA transcriptome analysis using the peripheral blood of patients developing RA-LPD to investigate the pathophysiology of RA-LPD.
Methods: Peripheral blood samples of the patients with RA-LPD were obtained between May 2013 and October 2018 from 30 hospitals in Japan. Whole transcriptome sequencing was performed using the Ion AmpliSeq Human Gene Expression Kit (Thermo Fisher Scientific) and an IonS5 XL sequencer (Thermo Fisher Scientific). The kit covers the expression levels of 20,802 human RefSeq genes (based on UCSC hg19). Differential gene expression analysis was performed by Transcriptome Analysis Console (TAC) software (ver. 4.0.3, Thermo Fisher Scientific) with |FC| > 2-fold difference and at p < 0.1. A Gene Set Enrichment Analysis (GSEA) was performed to identify pathways enriched in the Molecular Signatures Database (MSigDB) Hallmark gene set. A nominal p value of < 0.05 and an FDR (false discovery rate) q value of < 0.25 were considered statistically significant.
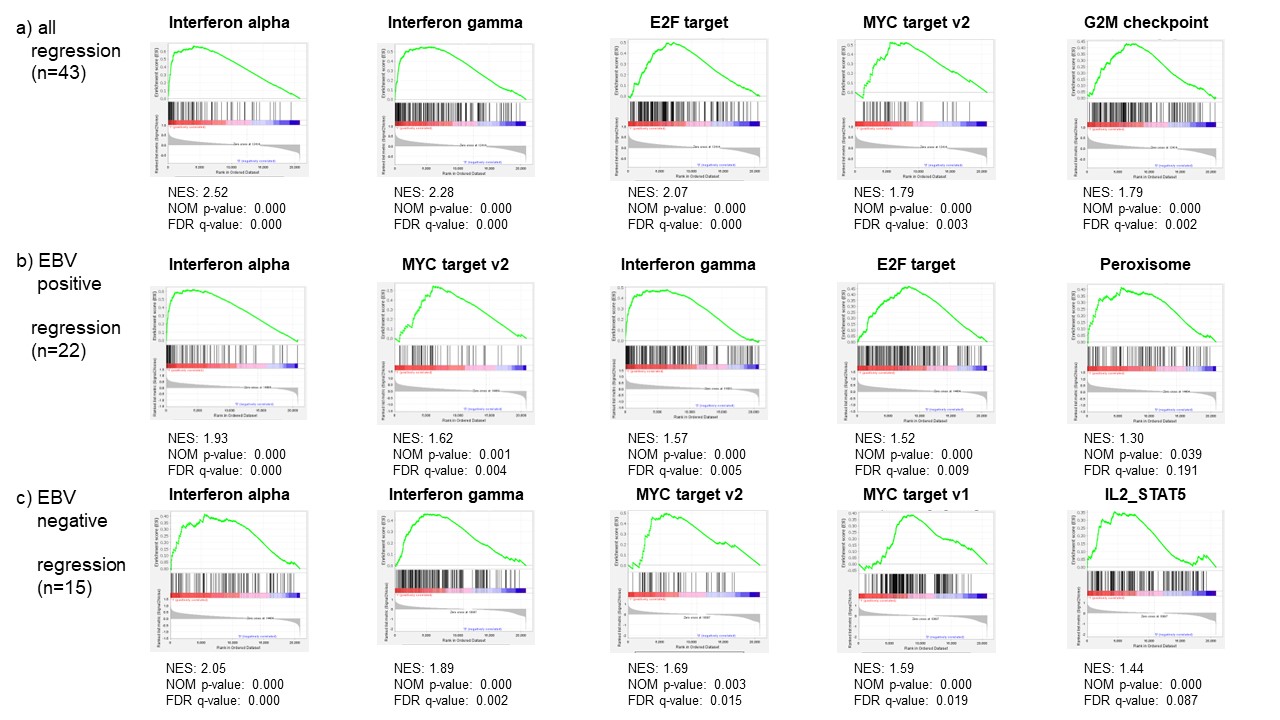
Results: In total, 211 cases (191 RA-LPD cases and 20 non-LPD RA cases) were analyzed. The flow diagram of the study is shown in Figure 1. We grouped diffuse large B-cell lymphoma (DLBCL) cases into two groups; regression cases (n = 43), which regressed spontaneously after stopping DMARDs and non-regression cases (n = 9), which did not regress. Gene expression levels were compared between the two groups. GSEA analysis showed that interferon alpha and gamma pathways were enhanced in regression cases (Figure 2a). To investigate mechanisms of regression in EBV-negative cases, we performed further analysis according to EBV infection status. (EBV positive, n= 22; EBV negative, n= 15). The interleukin-2-STAT5 (IL2-STAT5) pathway was enhanced in regression cases without EBV infection, which was suggested to be a unique mechanism for regression in EBV-negative RA-LPD (Figure 2b, c). The heatmap (Figure 3) shows that genes related to IL2-STAT5 were expressed highly in regression cases without EBV infection. Additionally, expression levels of some genes related to the IL2-STAT5 pathway were significantly different between regression and non-regression EBV-negative cases. This result suggests that these genes may serve as new liquid markers for regression in EBV-negative RA-LPD.
Conclusion: The current study showed that interferon-alpha and gamma pathways were upregulated in the regression cases with DLBCL of RA-LPD. Moreover, the study revealed a unique mechanism related to IL2-STAT5 for regression of EBV-negative cases. Our research further suggested new potential markers for regression in EBV-negative cases.
.jpg)
A. TSUJII: None; K. SAKAI: Chugai Pharmaceutical, 6, Life Technologies Japan, 6, Qiagen, 6, Takeda Pharmaceutical, 6; S. OHSHIMA: None; Y. SAEKI: None; M. YAGITA: Medical & Biological Laboratories, CO., Ltd, 9, ONK therapeutics, 9; T. MIYAMURA: None; M. Katayama: None; Y. HIRAMATSU: None; S. HIGA: None; F. HIRANO: None; K. ICHIKAWA: None; N. CHIBA: None; T. SUGIYAMA: None; A. IHATA: None; H. TSUTANI: None; K. TAKAHI: None; K. MIGITA: None; S. MORI: AbbVie GK, 6, 6, Asahikasei Pharma Corp, 6, Boehringer Ingelheim Japan, 6, Chugai Pharmaceutical Co. Ltd., 6, Chugai Pharmaceutical Co., Ltd., 6, Eli Lilly Japan K.K., 6, Janssen Pharmaceutical K.K., 6, Pfizer Japan Inc., 6, Taisho Pharma Co., Ltd., 6; N. YOSHIKAWA: None; A. UEDA: None; S. NAGAOKA: None; K. SETOGUCHI: None; S. SUGII: Eisai CO., Ltd., 6, MOCHIDA PHARMACEUTICAL CO>, 6; A. ABE: AbbVie/Abbott, 6, Bristol-Myers Squibb(BMS), 6; T. SUGAYA: None; H. SUGAHARA: None; S. TSUNODA: None; N. IIZUKA: None; R. YOSHIHARA: None; H. YABE: None; T. FUJISAKI: None; E. MORII: None; K. SAITO: None; K. Matsui: None; Y. TOMITA: None; H. FURUKAWA: None; S. Tohma: AbbVie/Abbott, 5, AsahiKASEI Co., Ltd., 6, Chudai Pharmaceutical Co., Ltd., 5, Mitsubishi Tanabe Pharma Corporation, 5, Pfizer Japan Inc., 6; K. NISHIO: Amgen, 6, AstraZeneca, 6, Bristol-Myers Squibb, 6, Chugai, 6, DAIICHI SANKYO, 6, Eli Lilly Japan, 5, 6, FUJIREBIO, 6, Guardant Health, 6, Hitachi, 5, Invitae Japan, 6, Janssen Pharmaceutical, 6, Merck Biopharma, 6, MSD, 6, Nichirei Biosciences, 5, Nippon Boehringer Ingelheim, 5, 6, Novartis Pharma, 6, Ono, 6, Otsuka Pharmaceutical, 5, 6, Pfizer, 6, SymBio Pharmaceuticalsっっv, 6, SYSMEX, 5, Takeda, 6, Yakult Honsha, 6; Y. HOSHIDA: None.
Background/Purpose: Lymphoproliferative disorders (LPD) in rheumatoid arthritis (RA) (RA-LPD) have unique pathophysiological features. More than half of the cases of RA-LPD undergo spontaneous regression after the discontinuation of disease modifying anti rheumatic drugs (DMARDs), whereas approximately 33% of these cases relapse. Epstein-Barr virus (EBV) infection is especially involved in the pathophysiology of RA-LPD. The current study was conducted with whole RNA transcriptome analysis using the peripheral blood of patients developing RA-LPD to investigate the pathophysiology of RA-LPD.
Methods: Peripheral blood samples of the patients with RA-LPD were obtained between May 2013 and October 2018 from 30 hospitals in Japan. Whole transcriptome sequencing was performed using the Ion AmpliSeq Human Gene Expression Kit (Thermo Fisher Scientific) and an IonS5 XL sequencer (Thermo Fisher Scientific). The kit covers the expression levels of 20,802 human RefSeq genes (based on UCSC hg19). Differential gene expression analysis was performed by Transcriptome Analysis Console (TAC) software (ver. 4.0.3, Thermo Fisher Scientific) with |FC| > 2-fold difference and at p < 0.1. A Gene Set Enrichment Analysis (GSEA) was performed to identify pathways enriched in the Molecular Signatures Database (MSigDB) Hallmark gene set. A nominal p value of < 0.05 and an FDR (false discovery rate) q value of < 0.25 were considered statistically significant.
Results: In total, 211 cases (191 RA-LPD cases and 20 non-LPD RA cases) were analyzed. The flow diagram of the study is shown in Figure 1. We grouped diffuse large B-cell lymphoma (DLBCL) cases into two groups; regression cases (n = 43), which regressed spontaneously after stopping DMARDs and non-regression cases (n = 9), which did not regress. Gene expression levels were compared between the two groups. GSEA analysis showed that interferon alpha and gamma pathways were enhanced in regression cases (Figure 2a). To investigate mechanisms of regression in EBV-negative cases, we performed further analysis according to EBV infection status. (EBV positive, n= 22; EBV negative, n= 15). The interleukin-2-STAT5 (IL2-STAT5) pathway was enhanced in regression cases without EBV infection, which was suggested to be a unique mechanism for regression in EBV-negative RA-LPD (Figure 2b, c). The heatmap (Figure 3) shows that genes related to IL2-STAT5 were expressed highly in regression cases without EBV infection. Additionally, expression levels of some genes related to the IL2-STAT5 pathway were significantly different between regression and non-regression EBV-negative cases. This result suggests that these genes may serve as new liquid markers for regression in EBV-negative RA-LPD.
Conclusion: The current study showed that interferon-alpha and gamma pathways were upregulated in the regression cases with DLBCL of RA-LPD. Moreover, the study revealed a unique mechanism related to IL2-STAT5 for regression of EBV-negative cases. Our research further suggested new potential markers for regression in EBV-negative cases.
.jpg)
Figure 1. Flow diagram of the study.
Cases analyzed in this study are written in red. RA-LPD: lymphoproliferative disorders in rheumatoid arthritis; DLBCL: diffuse large B-cell lymphoma; DMARD: Disease modifying anti rheumatic drug; EBV: Epstein-Barr virus.
Cases analyzed in this study are written in red. RA-LPD: lymphoproliferative disorders in rheumatoid arthritis; DLBCL: diffuse large B-cell lymphoma; DMARD: Disease modifying anti rheumatic drug; EBV: Epstein-Barr virus.

Figure 2. Gene set enrichment analysis (GSEA) in all regression cases (a), regression cases with Epstein-Barr virus (EBV) infection (b), and regression cases without EBV infection (c). Interferon alpha and gamma pathways were upregulated in all regression cases. Interleukin-2- STAT5 was a unique pathway in EBV negative cases.

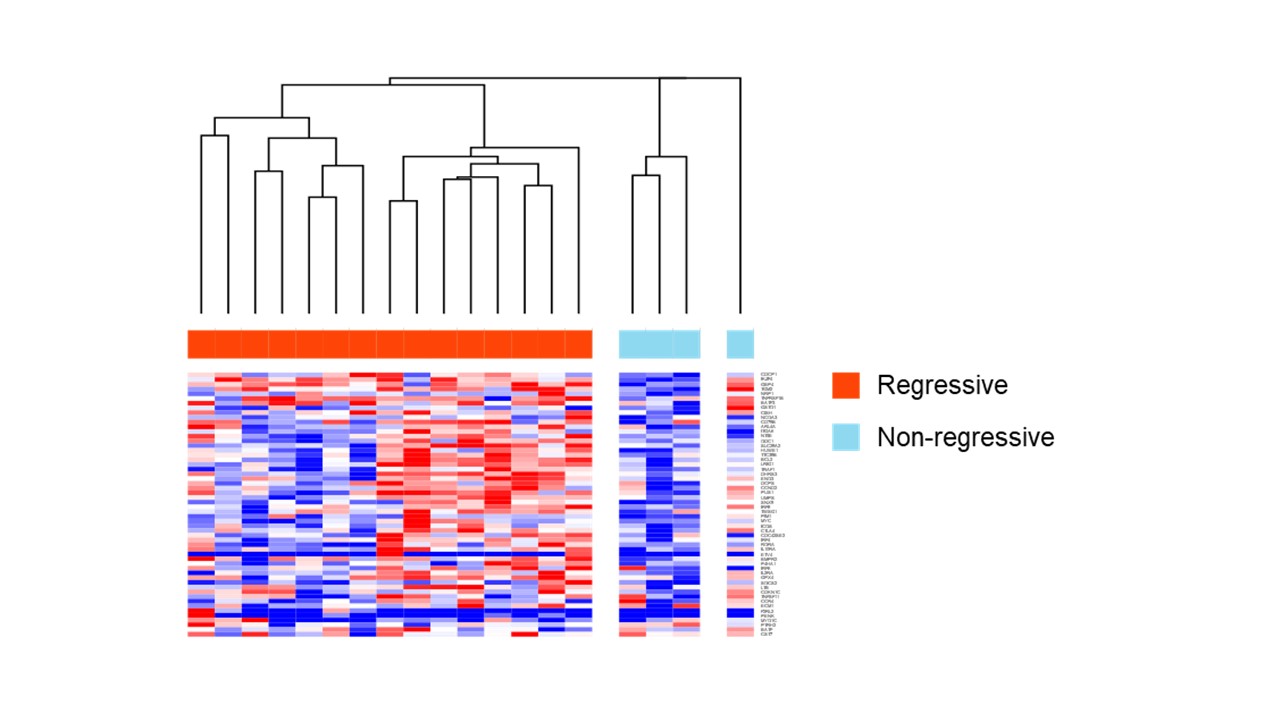
Figure 3. Heatmap showing gene expression related to interleukin 2-STAT5 pathway in cases without Epstein-Barr virus infection. They were highly expressed in regression cases.
A. TSUJII: None; K. SAKAI: Chugai Pharmaceutical, 6, Life Technologies Japan, 6, Qiagen, 6, Takeda Pharmaceutical, 6; S. OHSHIMA: None; Y. SAEKI: None; M. YAGITA: Medical & Biological Laboratories, CO., Ltd, 9, ONK therapeutics, 9; T. MIYAMURA: None; M. Katayama: None; Y. HIRAMATSU: None; S. HIGA: None; F. HIRANO: None; K. ICHIKAWA: None; N. CHIBA: None; T. SUGIYAMA: None; A. IHATA: None; H. TSUTANI: None; K. TAKAHI: None; K. MIGITA: None; S. MORI: AbbVie GK, 6, 6, Asahikasei Pharma Corp, 6, Boehringer Ingelheim Japan, 6, Chugai Pharmaceutical Co. Ltd., 6, Chugai Pharmaceutical Co., Ltd., 6, Eli Lilly Japan K.K., 6, Janssen Pharmaceutical K.K., 6, Pfizer Japan Inc., 6, Taisho Pharma Co., Ltd., 6; N. YOSHIKAWA: None; A. UEDA: None; S. NAGAOKA: None; K. SETOGUCHI: None; S. SUGII: Eisai CO., Ltd., 6, MOCHIDA PHARMACEUTICAL CO>, 6; A. ABE: AbbVie/Abbott, 6, Bristol-Myers Squibb(BMS), 6; T. SUGAYA: None; H. SUGAHARA: None; S. TSUNODA: None; N. IIZUKA: None; R. YOSHIHARA: None; H. YABE: None; T. FUJISAKI: None; E. MORII: None; K. SAITO: None; K. Matsui: None; Y. TOMITA: None; H. FURUKAWA: None; S. Tohma: AbbVie/Abbott, 5, AsahiKASEI Co., Ltd., 6, Chudai Pharmaceutical Co., Ltd., 5, Mitsubishi Tanabe Pharma Corporation, 5, Pfizer Japan Inc., 6; K. NISHIO: Amgen, 6, AstraZeneca, 6, Bristol-Myers Squibb, 6, Chugai, 6, DAIICHI SANKYO, 6, Eli Lilly Japan, 5, 6, FUJIREBIO, 6, Guardant Health, 6, Hitachi, 5, Invitae Japan, 6, Janssen Pharmaceutical, 6, Merck Biopharma, 6, MSD, 6, Nichirei Biosciences, 5, Nippon Boehringer Ingelheim, 5, 6, Novartis Pharma, 6, Ono, 6, Otsuka Pharmaceutical, 5, 6, Pfizer, 6, SymBio Pharmaceuticalsっっv, 6, SYSMEX, 5, Takeda, 6, Yakult Honsha, 6; Y. HOSHIDA: None.



