Poster Session A
Immunobiology
Session: (0066–0095) T Cell Biology & Targets in Autoimmune & Inflammatory Disease Poster
0075: Proteomic, Transcriptomic, and T-Cell Receptor (TCR) Profiling of Synovial Integrin-Expressing (InEx) T Cells in Axial Spondyloarthritis (axSpA)
Sunday, November 12, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall
- ZQ
Zoya Qaiyum, MSc
Krembil Research Institute
Toronto, ON, CanadaDisclosure information not submitted.
Abstract Poster Presenter(s)
Zoya Qaiyum1, Michael Tang2 and Robert Inman2, 1Krembil Research Institute, Toronto, ON, Canada, 2University Health Network, Toronto, ON, Canada
Background/Purpose: The strong clinical and genetic associations between axial spondyloarthritis (axSpA) and inflammatory bowel disease (IBD) underscore the pathogenic role of the gut-joint axis. The prevailing arthritogenic peptide hypothesis suggests that CD8+ T cells mount an immunological response upon recognition of a self or microbial arthritogenic peptide at inflammatory sites such as the gut. This could result in a clonal expansion of these cells, ultimately leading to their aberrant migration to other sites such as the joint. We previously identified a subpopulation of pathogenic mature CD8+ T cells in the axSpA synovial fluid (SF), expressing CD103+ and CD49a+ integrins (InEx cells). The expression of integrins on InEx cells implicates their role in aberrant migration between the gut-joint axis. Whether InEx cells recognize an arthritogenic peptide remains unclear. We hypothesize that InEx cells may incite and perpetuate chronic inflammation in axSpA. To this end, we characterized their T-cell receptor (TCR) repertoire and gene expression profile.
Methods: Patients fulfilling the ASAS classification for axSpA were recruited (n=4). Active joint effusion from these patients is reflective of active joint inflammation which coincided with high disease activity (BASDAI >4). We isolated peripheral blood mononuclear cells (PBMCs) and synovial fluid mononuclear cells (SFMCs), followed by FACS-sorting of mature CD8+ T cells and InEx cells. Subsequently, single-cell TCR (both α and β chains) and RNA sequencing were performed on these cell types. An HLA-B27+ reactive arthritis (ReA) sample were used as a comparison since it is the paradigm for antigen-driven inflammation initiated in the gut. Mature CD8+ T cells from ReA PBMCs and SFMCs were also FACS-sorted. These were all compared to HLA-B27+ healthy controls (n=4).
Results: The InEx TCR repertoire from axSpA patients was less diverse than mature CD8+ T cells from axSpA PBMCs or SFMCs, indicating a clonally expanded population. The TCR V/J pairings from the β chain in InEx cells confirm existing literature. Interestingly, their TRBV9/TRBJ2-3 usage was also found in mature CD8+ T cells from ReA SFMCs. Assessing the TCR V region from paired α/β chains revealed that the InEx cells shared similar usage with mature CD8+ T cells from ReA PBMCs or SFMCs: TRBV20-1/TRAV1-2, TRBV6-1/TRAV1-2, and TRBV6-2/TRAV1-2. Further, the CDR3 region of the β chain in InEx cells differed from mature CD8+ T cells from ReA SFMCs by only three amino acids. On a transcript level, InEx cells differed from ReA mature CD8+ T cells based on elevated genes such as CCL4L2, IFITM1, and IFITM2, and downregulated genes such as IL7R, GNLY, and KLRB1.
Conclusion: These observations suggest that the autoimmune mechanism employed by InEx cells could be mediated by HLA-B27+. This may enable them to incite inflammation similarly to ReA due to potential recognition of similar peptides, ultimately driving the selection of specific T cell clones. However, InEx cells' ability to perpetuate inflammation may be distinct from ReA due to a varied transcriptome. This has important implications for therapeutic designs attempting to target integrin blockade in axSpA.
Z. Qaiyum: None; M. Tang: None; R. Inman: AbbVie, 2, Eli Lilly, 2, Janssen, 2, Novartis, 2, Sandoz, 2.
Background/Purpose: The strong clinical and genetic associations between axial spondyloarthritis (axSpA) and inflammatory bowel disease (IBD) underscore the pathogenic role of the gut-joint axis. The prevailing arthritogenic peptide hypothesis suggests that CD8+ T cells mount an immunological response upon recognition of a self or microbial arthritogenic peptide at inflammatory sites such as the gut. This could result in a clonal expansion of these cells, ultimately leading to their aberrant migration to other sites such as the joint. We previously identified a subpopulation of pathogenic mature CD8+ T cells in the axSpA synovial fluid (SF), expressing CD103+ and CD49a+ integrins (InEx cells). The expression of integrins on InEx cells implicates their role in aberrant migration between the gut-joint axis. Whether InEx cells recognize an arthritogenic peptide remains unclear. We hypothesize that InEx cells may incite and perpetuate chronic inflammation in axSpA. To this end, we characterized their T-cell receptor (TCR) repertoire and gene expression profile.
Methods: Patients fulfilling the ASAS classification for axSpA were recruited (n=4). Active joint effusion from these patients is reflective of active joint inflammation which coincided with high disease activity (BASDAI >4). We isolated peripheral blood mononuclear cells (PBMCs) and synovial fluid mononuclear cells (SFMCs), followed by FACS-sorting of mature CD8+ T cells and InEx cells. Subsequently, single-cell TCR (both α and β chains) and RNA sequencing were performed on these cell types. An HLA-B27+ reactive arthritis (ReA) sample were used as a comparison since it is the paradigm for antigen-driven inflammation initiated in the gut. Mature CD8+ T cells from ReA PBMCs and SFMCs were also FACS-sorted. These were all compared to HLA-B27+ healthy controls (n=4).
Results: The InEx TCR repertoire from axSpA patients was less diverse than mature CD8+ T cells from axSpA PBMCs or SFMCs, indicating a clonally expanded population. The TCR V/J pairings from the β chain in InEx cells confirm existing literature. Interestingly, their TRBV9/TRBJ2-3 usage was also found in mature CD8+ T cells from ReA SFMCs. Assessing the TCR V region from paired α/β chains revealed that the InEx cells shared similar usage with mature CD8+ T cells from ReA PBMCs or SFMCs: TRBV20-1/TRAV1-2, TRBV6-1/TRAV1-2, and TRBV6-2/TRAV1-2. Further, the CDR3 region of the β chain in InEx cells differed from mature CD8+ T cells from ReA SFMCs by only three amino acids. On a transcript level, InEx cells differed from ReA mature CD8+ T cells based on elevated genes such as CCL4L2, IFITM1, and IFITM2, and downregulated genes such as IL7R, GNLY, and KLRB1.
Conclusion: These observations suggest that the autoimmune mechanism employed by InEx cells could be mediated by HLA-B27+. This may enable them to incite inflammation similarly to ReA due to potential recognition of similar peptides, ultimately driving the selection of specific T cell clones. However, InEx cells' ability to perpetuate inflammation may be distinct from ReA due to a varied transcriptome. This has important implications for therapeutic designs attempting to target integrin blockade in axSpA.

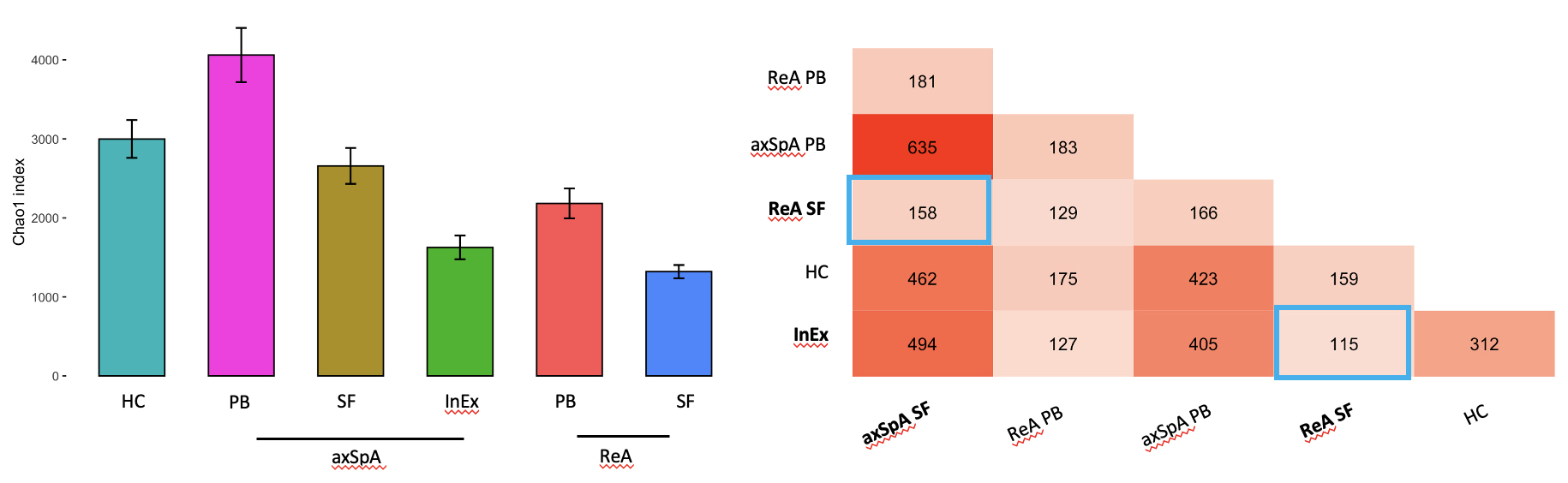
Left panel displaying the Chao1 index, a measure of TCR repertoire diversity, stratified by sample type. Right panel displaying the overlapping alpha and beta TCR chain gene usage of the V region. Highlighted in blue boxes are features similar between InEx and ReA SF and axSpA SF (mature CD8+ T cells) and ReA SF. HC = healthy controls, PB = peripheral blood, SF = synovial fluid.

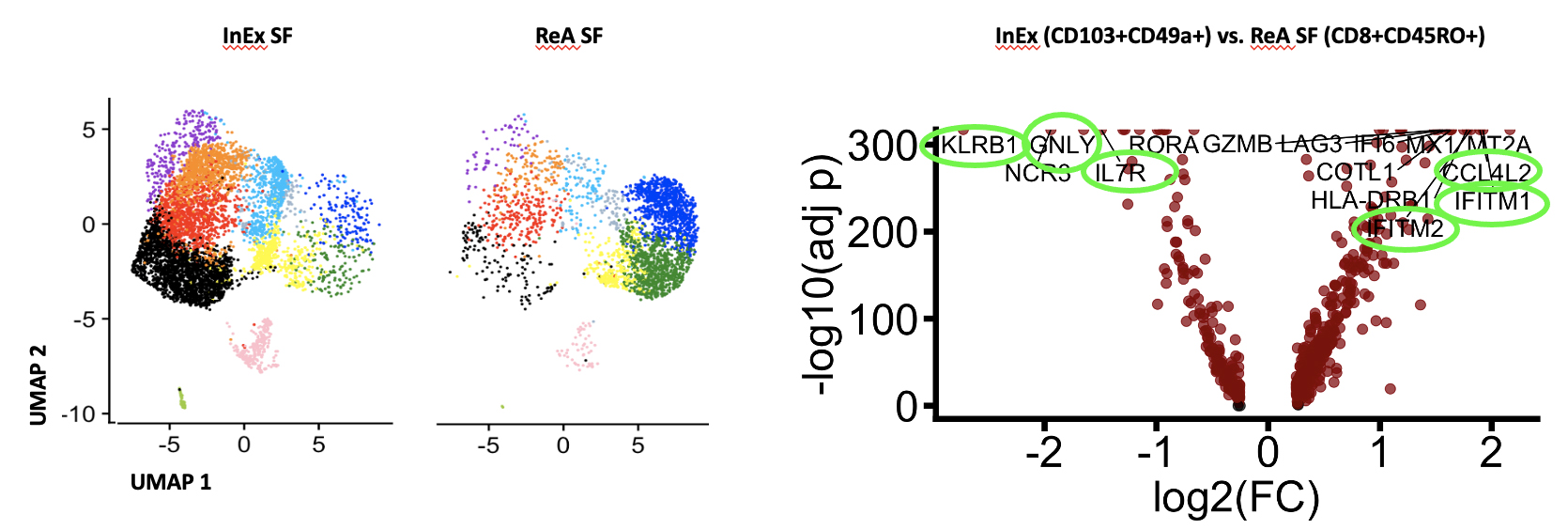
Left panel displaying transcriptome profile of InEx vs mature CD8+ ReA SFMCs on UMAP plots. Genetic profile of InEx cells cluster differently to ReA SF. Right panel displaying differently expressed genes between the two groups, with most informative genes circled in green. UMAP = Uniform Manifold Approximation and Projection. SF = synovial fluid.
Z. Qaiyum: None; M. Tang: None; R. Inman: AbbVie, 2, Eli Lilly, 2, Janssen, 2, Novartis, 2, Sandoz, 2.



