Abstract Session
Systemic lupus erythematosus (SLE)
Session: Abstracts: SLE – Diagnosis, Manifestations, & Outcomes I: Biomarkers (0847–0852)
0849: Multiplex Profiling and Machine Learning Reveal Distinct Signatures of Circulating Cytokines Associated with Autoantibody Profiles and Disease Severity in Systemic Lupus Erythematosus
Sunday, November 12, 2023
4:30 PM - 4:40 PM PT
Location: Exhibit Hall A-B
- SP
Sarit Pattanaik, MD, DM
SCB Medical College and Hospital, Cuttack, Odisha
Cuttack, Odisha, IndiaDisclosure information not submitted.
Presenting Author(s)
Sarit Pattanaik1, Ratnadeep Mukherjee2, Rina Tripathy3, Birendra Prusty4, Balachandran Ravindran5 and Bidyut Das6, 1SCB Medical College and Hospital, Cuttack, Odisha, Cuttack, India, 2Norwegian Institute of Public Health, Oslo, Norway, 3Sardar Vallabhbhai Patel Post Graduate Institute of Pediatrics, Cuttack, India, 4Fisheries & Animal Resources Development Department, Phulbani, India, 5Institute of Life Sciences, Bhubaneswar, India, 6SCB medical college, Cuttack, India
Background/Purpose: SLE is one of the leading causes of death in young females suffering from autoimmune disorders. Nephritis, afflicts 60-70% of patients which contribute significantly to morbidity and mortality despite therapeutic advances. Disrupted cytokine networks and autoantibodies play an important role in disease pathogenesis. However, conflicting reports and non-reproducibility have hindered progress with regards to translational potential of cytokines. This study attempts to address the existing knowledge gap using a multiplex cytokine assay and machine learning algorithms.
Methods: 69 SLE patients fulfilling SLICC criteria were recruited. Baseline characteristics along with disease activity was recorded for all patients. Cytokines were measured by a commercially available multiplex cytokine kit (Millipore). Fluorescence values were logarithmically transformed before analysis. To visualize relationships between cytokines, Network graphs were constructed based on Spearman correlation values. For stratifying individuals based on cytokine profiles, we used Sparse Partial Least Squares Discriminant Analysis (sPLS-DA) followed by factor loadings plots to assess which cytokines contribute most to the observed differences between groups. Patterns of co-occurrence of autoantibodies within the SLE patients were identified by the K-Modes algorithm and the patients were visualized on a dimensionally reduced plot obtained by constructing a 2-dimensional representation by multiple correspondence analysis (MCA).
Results: We observed a positive association between actual disease activity scores (SLEDAI) and predicted scores from a partial least squares regression (PLSR) analysis of multivariate cytokine response data (Figure 1a). A study of the first two PLSR components revealed the SLEDAI scores to be positively associated with the first component (LV1) and negatively associated with the second component (LV2) (Figure 1b). Analysis of cytokine contributing towards the first two PLSR components revealed MIP-1α to have a strong positive influence towards component 1 and a negative influence on component 2, indicating a strong association with increased disease severity (Figure 1c and d). K-Modes clustering analysis identified 4 distinct clusters of patients with specific autoantibodies (Figure 2a and b), with clusters 2 and 3 being the most well-separated. Furthermore, we also observed striking differences in distributions of lupus nephritis between the clusters, with all patients in cluster 3 presenting with nephritis (Figure 2c). Finally, our results also demonstrate unique cytokine signatures associated with autoantibody profiles, with patients in cluster 3 showing significantly lower levels of cytokines responsible for modulating inflammation and maintaining cellular homeostasis (Figure 3), an observation that is in line with the clinical feature of nephritis.
Conclusion: Cytokine response can predict disease activity. Nephritis is associated with specific autoantibody profiles and cytokine signatures.
.jpg)
.jpg)
S. Pattanaik: None; R. Mukherjee: None; R. Tripathy: None; B. Prusty: None; B. Ravindran: None; B. Das: None.
Background/Purpose: SLE is one of the leading causes of death in young females suffering from autoimmune disorders. Nephritis, afflicts 60-70% of patients which contribute significantly to morbidity and mortality despite therapeutic advances. Disrupted cytokine networks and autoantibodies play an important role in disease pathogenesis. However, conflicting reports and non-reproducibility have hindered progress with regards to translational potential of cytokines. This study attempts to address the existing knowledge gap using a multiplex cytokine assay and machine learning algorithms.
Methods: 69 SLE patients fulfilling SLICC criteria were recruited. Baseline characteristics along with disease activity was recorded for all patients. Cytokines were measured by a commercially available multiplex cytokine kit (Millipore). Fluorescence values were logarithmically transformed before analysis. To visualize relationships between cytokines, Network graphs were constructed based on Spearman correlation values. For stratifying individuals based on cytokine profiles, we used Sparse Partial Least Squares Discriminant Analysis (sPLS-DA) followed by factor loadings plots to assess which cytokines contribute most to the observed differences between groups. Patterns of co-occurrence of autoantibodies within the SLE patients were identified by the K-Modes algorithm and the patients were visualized on a dimensionally reduced plot obtained by constructing a 2-dimensional representation by multiple correspondence analysis (MCA).
Results: We observed a positive association between actual disease activity scores (SLEDAI) and predicted scores from a partial least squares regression (PLSR) analysis of multivariate cytokine response data (Figure 1a). A study of the first two PLSR components revealed the SLEDAI scores to be positively associated with the first component (LV1) and negatively associated with the second component (LV2) (Figure 1b). Analysis of cytokine contributing towards the first two PLSR components revealed MIP-1α to have a strong positive influence towards component 1 and a negative influence on component 2, indicating a strong association with increased disease severity (Figure 1c and d). K-Modes clustering analysis identified 4 distinct clusters of patients with specific autoantibodies (Figure 2a and b), with clusters 2 and 3 being the most well-separated. Furthermore, we also observed striking differences in distributions of lupus nephritis between the clusters, with all patients in cluster 3 presenting with nephritis (Figure 2c). Finally, our results also demonstrate unique cytokine signatures associated with autoantibody profiles, with patients in cluster 3 showing significantly lower levels of cytokines responsible for modulating inflammation and maintaining cellular homeostasis (Figure 3), an observation that is in line with the clinical feature of nephritis.
Conclusion: Cytokine response can predict disease activity. Nephritis is associated with specific autoantibody profiles and cytokine signatures.
.jpg)
Figure 1: Combined cytokine response in SLE is predictive of disease severity. (A) Correlation between observed vs. predicted SLEDAI scores in patients with SLE. The SLEDAI scores were predicted from a PLSR model with cytokines. (B) Correlation of SLEDAI scores with the first two components of the PLSR model. (C) Cytokine contributions towards the first PLSR component (LV1: Latent variable 1). (D) Cytokine contributions towards the second PLSR component (LV2: Latent variable 2).
.jpg)
Figure 2: SLE patients have distinct combinations of autoantibody profiles that associate with lupus nephritis. (A) K Modes clustering algorithm identifies 4 distinct clusters of patients based on combined autoantibody profiles. The patients are then visualized on a 2D representation of the multidimensional data by multiple correspondence analysis (MCA). (B) Factor loadings plot demonstrating the relative contributions of the individual autoantibodies to the separation observed in (A). (C) Counts of patients presenting with or without lupus nephritis with the corresponding autoantibody clusters.

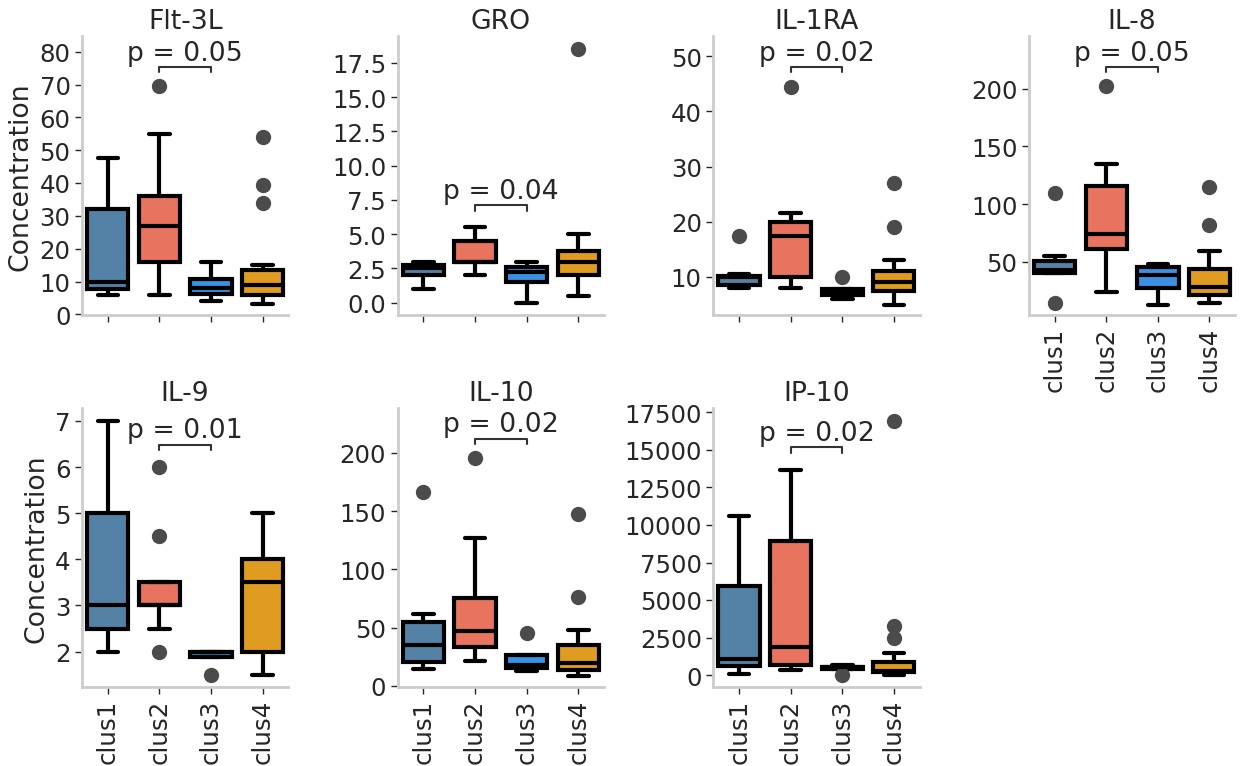
Figure 3: Cytokine profiles associated with autoantibody clusters. Comparison of cytokine levels between Clusters 2 and 3 was done by a Mann-Whitney U test, corrected for multiple tests by the Bonferroni method.
S. Pattanaik: None; R. Mukherjee: None; R. Tripathy: None; B. Prusty: None; B. Ravindran: None; B. Das: None.




