Poster Session C
Spondyloarthritis (SpA) including psoriatic arthritis (PsA)
Session: (1776–1795) Spondyloarthritis Including Psoriatic Arthritis – Basic Science Poster
1790: Expansion of CD8+ TCRVβ9+ T Cells in the Peripheral Blood of HLA-B27+ Patients with Axial Spondyloarthritis
Tuesday, November 14, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall
- CB
Caroline Bauchiero, MS (she/her/hers)
Brigham and Women's Hospital
Brookline, MA, United StatesDisclosure information not submitted.
Abstract Poster Presenter(s)
Caroline Bauchiero1, Micah Lefton1, Stephanie Sinnappan2, Jeffrey Sparks3 and Joerg Ermann4, 1Brigham and Women's Hospital, Boston, MA, 2Columbia University, New York, NY, 3Division of Rheumatology, Inflammation, and Immunity, Brigham and Women’s Hospital and Harvard Medical School, Boston, MA, 4Division of Rheumatology, Inflammation and Immunity, Brigham and Women's Hospital and Harvard Medical School, Boston, MA
Background/Purpose: Studies in human HLA-B27 transgenic rats largely discredited the idea that CD8+ T cells are key drivers of disease in axial spondyloarthritis (axSpA). However, recent studies in humans have revived interest in the role of CD8+ T cells in axSpA including the description of relevant CD8+ T cell subsets in a number of rheumatic disease settings (CD103+CD49a+ InEx cells, regulatory CD8+KIR+ T cells, Granzyme B and K expressing CD8+ T cell subsets) and the demonstration of expanded TCR clonotypes in the joints and eyes of HLA-B27+ SpA patients. In this study, we used state-of-the-art flow cytometry to search for diagnostic cellular biomarkers in axSpA reporting results for CD8+ T cells.
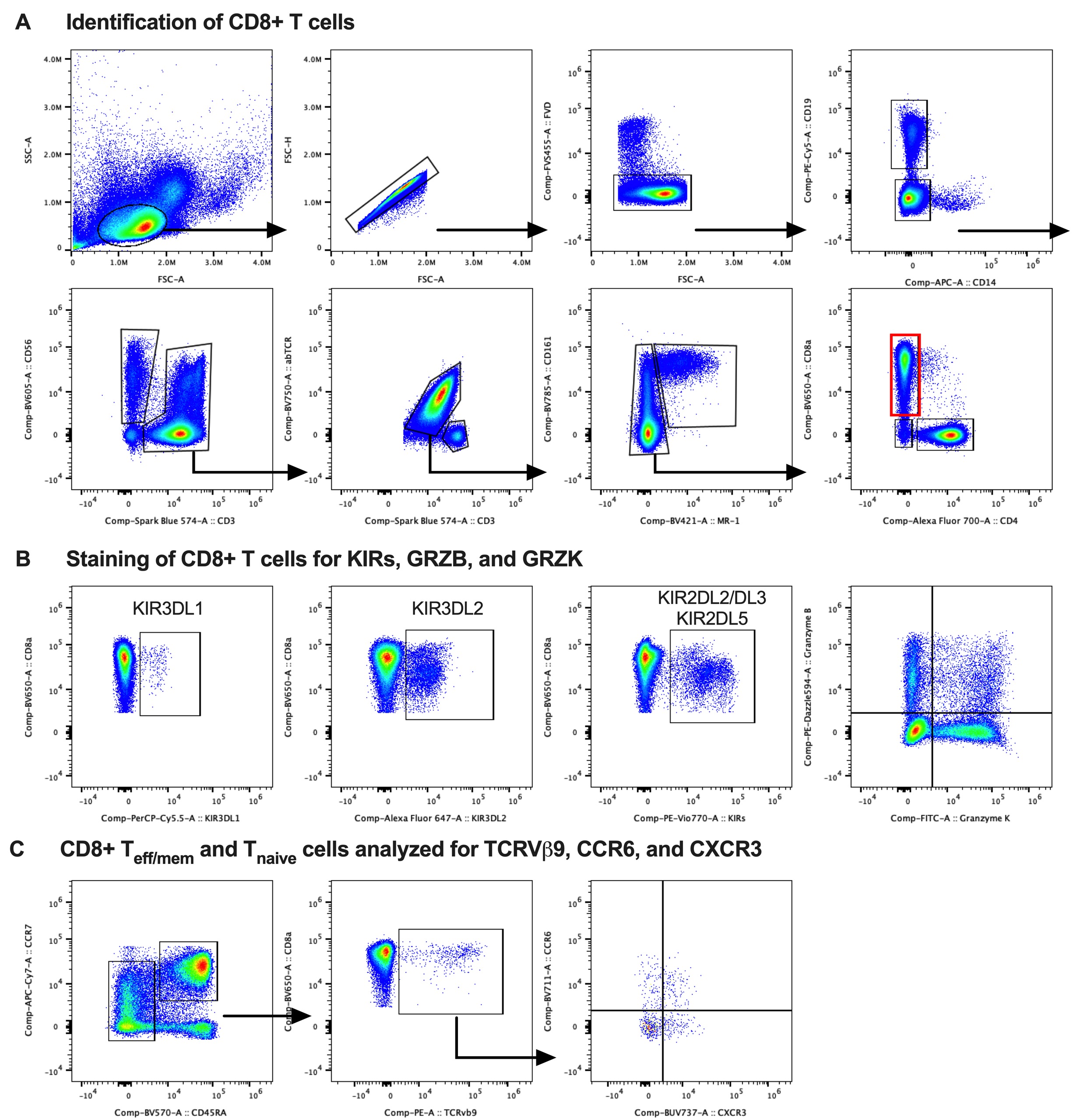
Methods: Study subjects were recruited from our hospital's Orthopedic and Arthritis Center. Four groups were compared: healthy controls (HC), patients with chronic low back pain without axSpA (cLBP), axSpA patients not on a biologic (axSpA/-), and axSpA patients treated with a TNF inhibitor (axSpA/TNFi). Groups (n=22-23 each) were matched for age, sex and genotyped for HLA-B27. All axSpA patients fulfilled modified New York criteria for ankylosing spondylitis or the 2009 ASAS criteria for axSpA. PBMCs were analyzed in two batches on a CYTEK Aurora spectral flow cytometer. Our customized 30-parameter staining panel included major lineage markers, lymphocyte differentiation and activation markers and functional markers for cytotoxicity, homing, and cytokine production potential. We included antibodies for KIR3DL1 (DX9), KIR3DL2 (DX31), KIR2DL2/L3 (DX27) and KIR2DL5 (UP-R1) as well as TCRVβ9+ (MKB1). Cells were further stained intracellularly for GRZB (GB11) and GRZK (GM26E7). FlowJo was used for quality control and analysis (see gating strategy in Fig. 1).
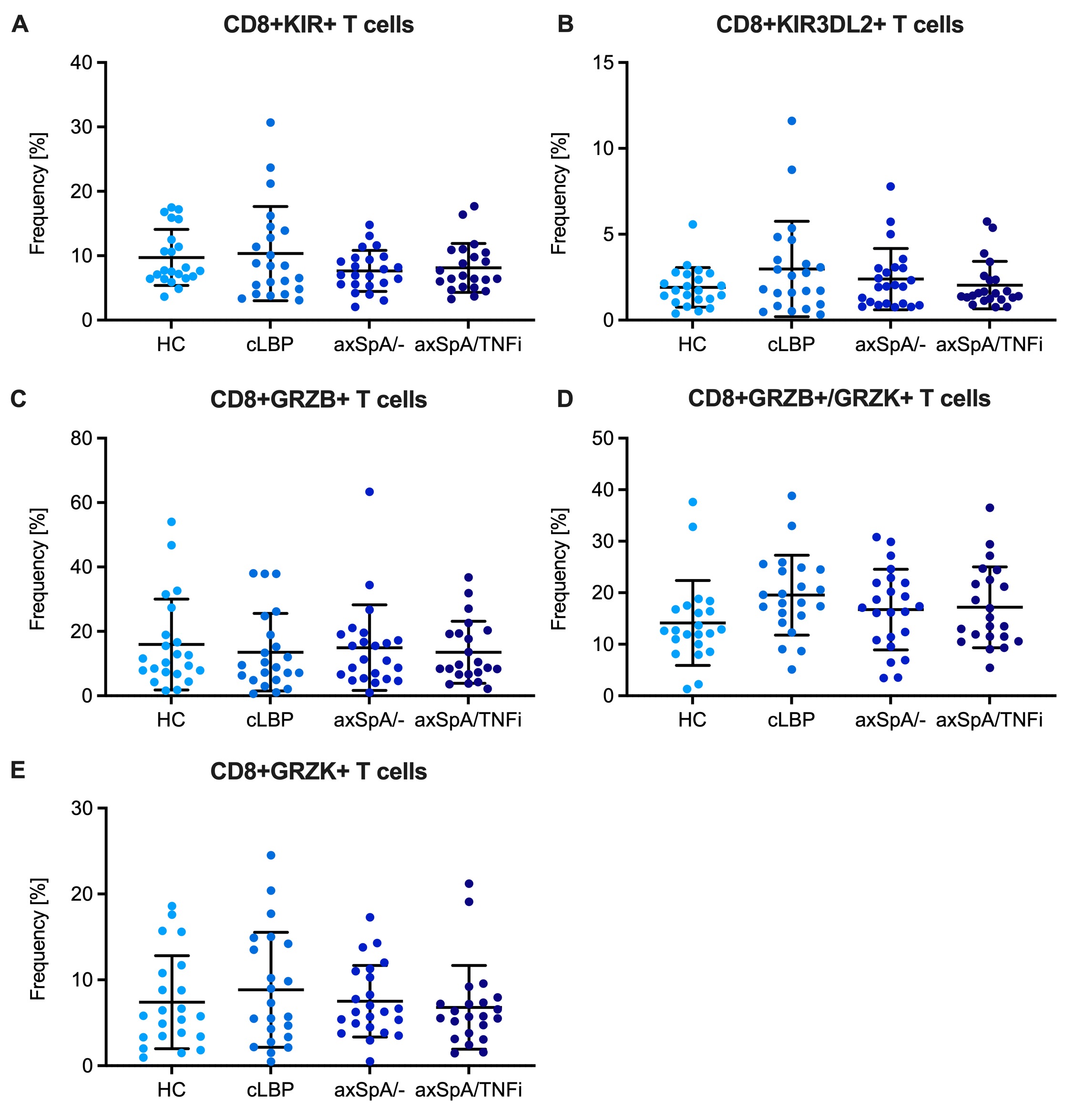
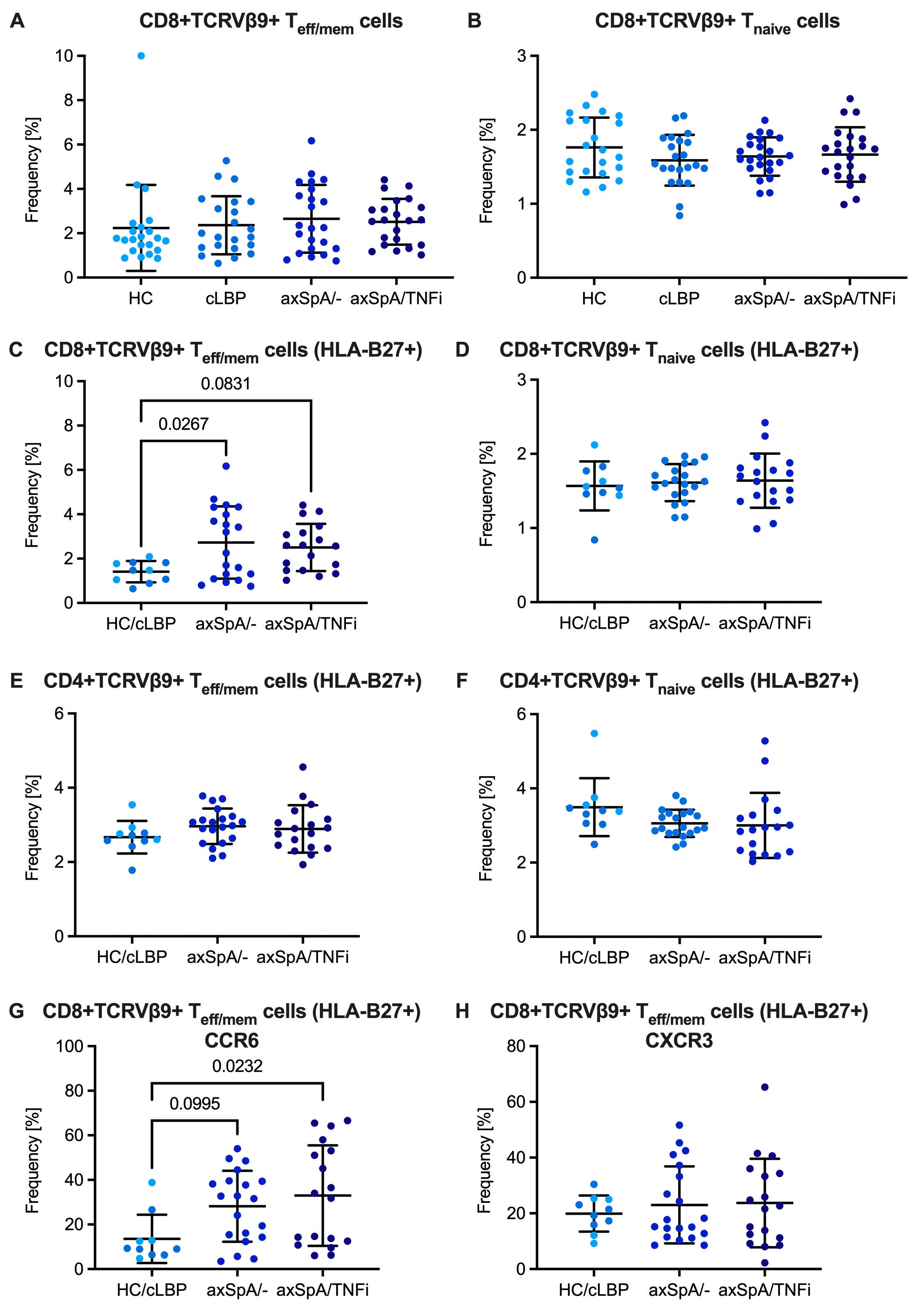
Results: There were no differences in the frequency of CD8+ T cells expressing any KIR or CD8+ T cells expressing the HLA-B27 binding KIR3DL2 between HC, cLBP, axSpA/- and axSpA/TNFi subjects (Fig. 2A,B). There were also no differences in the frequency of CD8+ T cell subsets distinguished by expression of GRZB and GRZK (Fig. 2C-E). About 2% of CD8+CD45RAloTeff/mem and CD8+CD45RAhiCCR7+ Tnaive cells were TCRVβ9 positive without differences between groups. However, when only HLA-B27+ individuals were analyzed, we noticed an expansion of TCRVβ9+ cells amongst CD8+ Teff/mem cells in axSpA/- and axSpA/TNFi subjects, which was not seen in CD8+ Tnaïve cells or in the respective CD4+ T cell subsets (Fig. 3). Compared with control subjects, CD8+TCRVβ9+ Teff/mem cells in axSpA patients showed increased expression of CCR6 suggesting a Tc17 phenotype.
Conclusion: The observed expansion of CD8+Vβ9+ Teff/mem cells in the peripheral blood of HLA-B27+ patients with axSpA is consistent with the previously reported expansion of TRBV9–CDR3–TRBJ2.3 clonotypes in tissues and supports the peptide hypothesis for the HLA-B27 association with axSpA. The in-depth characterization of the CD8+TCRVβ9+ Teff/mem population using additional markers included in the staining panel is ongoing.
C. Bauchiero: None; M. Lefton: None; S. Sinnappan: None; J. Sparks: AbbVie, 2, Amgen, 2, Boehringer Ingelheim, 2, Bristol-Myers Squibb, 2, 5, Gilead, 2, Inova Diagnostics, 2, Janssen, 2, Optum, 2, Pfizer, 2, ReCor, 2; J. Ermann: AbbVie, 2, 5, Boehringer Ingelheim, 5, Janssen, 2, Novartis, 2, 5, Pfizer, 2, 5, Takeda, 2, UCB Pharma, 2.
Background/Purpose: Studies in human HLA-B27 transgenic rats largely discredited the idea that CD8+ T cells are key drivers of disease in axial spondyloarthritis (axSpA). However, recent studies in humans have revived interest in the role of CD8+ T cells in axSpA including the description of relevant CD8+ T cell subsets in a number of rheumatic disease settings (CD103+CD49a+ InEx cells, regulatory CD8+KIR+ T cells, Granzyme B and K expressing CD8+ T cell subsets) and the demonstration of expanded TCR clonotypes in the joints and eyes of HLA-B27+ SpA patients. In this study, we used state-of-the-art flow cytometry to search for diagnostic cellular biomarkers in axSpA reporting results for CD8+ T cells.
Methods: Study subjects were recruited from our hospital's Orthopedic and Arthritis Center. Four groups were compared: healthy controls (HC), patients with chronic low back pain without axSpA (cLBP), axSpA patients not on a biologic (axSpA/-), and axSpA patients treated with a TNF inhibitor (axSpA/TNFi). Groups (n=22-23 each) were matched for age, sex and genotyped for HLA-B27. All axSpA patients fulfilled modified New York criteria for ankylosing spondylitis or the 2009 ASAS criteria for axSpA. PBMCs were analyzed in two batches on a CYTEK Aurora spectral flow cytometer. Our customized 30-parameter staining panel included major lineage markers, lymphocyte differentiation and activation markers and functional markers for cytotoxicity, homing, and cytokine production potential. We included antibodies for KIR3DL1 (DX9), KIR3DL2 (DX31), KIR2DL2/L3 (DX27) and KIR2DL5 (UP-R1) as well as TCRVβ9+ (MKB1). Cells were further stained intracellularly for GRZB (GB11) and GRZK (GM26E7). FlowJo was used for quality control and analysis (see gating strategy in Fig. 1).
Results: There were no differences in the frequency of CD8+ T cells expressing any KIR or CD8+ T cells expressing the HLA-B27 binding KIR3DL2 between HC, cLBP, axSpA/- and axSpA/TNFi subjects (Fig. 2A,B). There were also no differences in the frequency of CD8+ T cell subsets distinguished by expression of GRZB and GRZK (Fig. 2C-E). About 2% of CD8+CD45RAloTeff/mem and CD8+CD45RAhiCCR7+ Tnaive cells were TCRVβ9 positive without differences between groups. However, when only HLA-B27+ individuals were analyzed, we noticed an expansion of TCRVβ9+ cells amongst CD8+ Teff/mem cells in axSpA/- and axSpA/TNFi subjects, which was not seen in CD8+ Tnaïve cells or in the respective CD4+ T cell subsets (Fig. 3). Compared with control subjects, CD8+TCRVβ9+ Teff/mem cells in axSpA patients showed increased expression of CCR6 suggesting a Tc17 phenotype.
Conclusion: The observed expansion of CD8+Vβ9+ Teff/mem cells in the peripheral blood of HLA-B27+ patients with axSpA is consistent with the previously reported expansion of TRBV9–CDR3–TRBJ2.3 clonotypes in tissues and supports the peptide hypothesis for the HLA-B27 association with axSpA. The in-depth characterization of the CD8+TCRVβ9+ Teff/mem population using additional markers included in the staining panel is ongoing.

Gating strategy. (A) Identification of CD8+ T cells by hierarchical gating. (B) Staining of CD8+ T cells for expression of KIR3DL1, KIR3DL2, KIR2DL2/L3+KIR2DL5, and GRZB vs. GRZK. CD8+ T cells expressing any KIR were identified by Boolean gating (logic OR gate) in FlowJo. (C) CD45RA and CCR7 were used to identify CD8+ Teff/mem and Tnaive cells, which were then analyzed for expression of TCRVb9, CCR6 and CXCR3

(A) Expression of KIRs on total CD8+ T cells. Depicted is the fraction of CD8+ T cells that express any of the following KIRs: KIR3DL1, KIR3DL2, KIR2DL2/L3, KIR2DL5 (B) Fraction of KIR3DL2+ CD8+ T cells. (C-E) Fraction of CD8+ T cells expressing GRZB alone, GRZB and GRZK, or GRZK alone. Bars represent means and SD deviation. There were no significant differences between groups by one-way ANOVA.

(A, B) Fraction of TCRVb9+ cells amongst CD8+ Teff/mem cells and CD8+ Tnaive cells, respectively. (C, D) Fraction of TCRVb9+ cells amongst CD8+ Teff/mem cells and CD8+ Tnaive cells in HLA-B27+ individuals. HC (n=4) and cLBP (n=6) were pooled. (E, F) Fraction of TCRVb9+ cells amongst CD4+ Teff/mem cells and CD4+ Tnaive cells in HLA-B27+ individuals. (G, H) Fraction of CCR6+ and CXCR3+ cells amongst CD8+TCRVb9+ Teff/mem cells in HLA-B27+ individuals. Bars represent means and SD deviation. P values were calculated by one-way ANOVA with post hoc Tukey’s.
C. Bauchiero: None; M. Lefton: None; S. Sinnappan: None; J. Sparks: AbbVie, 2, Amgen, 2, Boehringer Ingelheim, 2, Bristol-Myers Squibb, 2, 5, Gilead, 2, Inova Diagnostics, 2, Janssen, 2, Optum, 2, Pfizer, 2, ReCor, 2; J. Ermann: AbbVie, 2, 5, Boehringer Ingelheim, 5, Janssen, 2, Novartis, 2, 5, Pfizer, 2, 5, Takeda, 2, UCB Pharma, 2.



