Poster Session A
Genetics, genomics and proteomics
Session: (0013–0039.5) Genetics, Genomics & Proteomics Poster
0031: A Systematic Approach for Identifying Causal Variants and Their Target Genes on JIA Risk Haplotypes
Sunday, November 12, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall
- KJ
Kaiyu Jiang, PhD
University at Buffalo
Buffalo, NY, United StatesDisclosure information not submitted.
Abstract Poster Presenter(s)
Kaiyu Jiang1, tao liu2, Ryan Tewhey3, Susan Kales3, Yungki Park4 and James N Jarvis5, 1University at Buffalo, Buffalo, NY, 2Roswell Park Cancer Institute, Buffalo, NY, 3Jackson Laboratories, Bar Harbor, ME, 4University at Buffalo Jacobs School of Medicine & Biomedical Sciences, Buffalo, NY, 5University at Buffalo Jacobs School of Medicine, Buffalo, NY
Background/Purpose: GWAS have identified multiple genetic regions that confer risk for juvenile idiopathic arthritis (JIA).However, identifying the single nucleotide polymorphisms (SNPs) that drive disease risk has been impeded by the fact that the SNPs used to identify risk loci are in linkage disequilibrium (LD) with hundreds of other SNPs. Since the causal SNPs remain unknown, it is difficult to identify target genes, and thus use genetic information to elucidate disease biology and inform patient care. We have therefore developed a functional genomics pipeline to identify causal variants and their target genes on JIA risk haplotypes.
Methods: We designed 180 bp oligonucleotides (“oligos”) representing 7,312 SNPs in LD with the tag SNPs on JIA risk loci (r2 = 0.80).Oligos were bar coded and cloned into a green fluorescence protein (GFP)-carrying plasmid vector behind a minimal promoter. We transfected our oligo libraries into myeloid K562 cells to conduct a massively parallel reporter assay (MPRA) in which GFP expression for each of the 7,312 SNPs was compared to the common allele. We then used 3D chromatin data to identify the chromatin loops harboring the enhancers carrying SNPs that screen positive on MPRA. We used K562 cells stably transfected with the epigenome editing enzyme dCas-KRAB and gRNAs across the enhancer regions in the TRAF1 and LNPEP/ERAP2 loci, inducing dCas-KRAB with doxycycline and using qrtPCR to query expression levels of genes within the same chromatin loops.
Results: We identified n=44 SNPs that showed a significant difference in gene expression (FC > 2.0, FDR = 0.05) when compared to the common allele. After stimulating K562 cells with IFNγ (250 ng/ml), we identified an additional 42 SNPs that showed significant effects on gene expression that were not identified in unstimulated cells. In many cases, we identified multiple alleles on the same haplotype, although these alleles were not necessarily within the same functional element (e.g., intergenic vs intronic enhancers). Ablating the intergenic enhancer harboring rs10985080 on the TRAF1 haplotype using CRISPRi significantly reduced the expression of PHF1, but not C5, which is in the same chromatin loop. Using a similar approach for an intergenic enhancer in the LNPEP/ERAP2 locus, we found reduced expression of both LNPEP and ERAP2 but not CAST.
Conclusion: We demonstrate proof-of concept for identifying causal variants on JIA risk haplotypes based on observed functional properties that distinguish them from the SNPs in which they are in LD. We also demonstrate that, once these variants are identified, target genes can be quickly identified using publicly available 3D chromatin data and additional functional assays such as CRISPRi.
K. Jiang: None; t. liu: None; R. Tewhey: None; S. Kales: None; Y. Park: None; J. Jarvis: None.
Background/Purpose: GWAS have identified multiple genetic regions that confer risk for juvenile idiopathic arthritis (JIA).However, identifying the single nucleotide polymorphisms (SNPs) that drive disease risk has been impeded by the fact that the SNPs used to identify risk loci are in linkage disequilibrium (LD) with hundreds of other SNPs. Since the causal SNPs remain unknown, it is difficult to identify target genes, and thus use genetic information to elucidate disease biology and inform patient care. We have therefore developed a functional genomics pipeline to identify causal variants and their target genes on JIA risk haplotypes.
Methods: We designed 180 bp oligonucleotides (“oligos”) representing 7,312 SNPs in LD with the tag SNPs on JIA risk loci (r2 = 0.80).Oligos were bar coded and cloned into a green fluorescence protein (GFP)-carrying plasmid vector behind a minimal promoter. We transfected our oligo libraries into myeloid K562 cells to conduct a massively parallel reporter assay (MPRA) in which GFP expression for each of the 7,312 SNPs was compared to the common allele. We then used 3D chromatin data to identify the chromatin loops harboring the enhancers carrying SNPs that screen positive on MPRA. We used K562 cells stably transfected with the epigenome editing enzyme dCas-KRAB and gRNAs across the enhancer regions in the TRAF1 and LNPEP/ERAP2 loci, inducing dCas-KRAB with doxycycline and using qrtPCR to query expression levels of genes within the same chromatin loops.
Results: We identified n=44 SNPs that showed a significant difference in gene expression (FC > 2.0, FDR = 0.05) when compared to the common allele. After stimulating K562 cells with IFNγ (250 ng/ml), we identified an additional 42 SNPs that showed significant effects on gene expression that were not identified in unstimulated cells. In many cases, we identified multiple alleles on the same haplotype, although these alleles were not necessarily within the same functional element (e.g., intergenic vs intronic enhancers). Ablating the intergenic enhancer harboring rs10985080 on the TRAF1 haplotype using CRISPRi significantly reduced the expression of PHF1, but not C5, which is in the same chromatin loop. Using a similar approach for an intergenic enhancer in the LNPEP/ERAP2 locus, we found reduced expression of both LNPEP and ERAP2 but not CAST.
Conclusion: We demonstrate proof-of concept for identifying causal variants on JIA risk haplotypes based on observed functional properties that distinguish them from the SNPs in which they are in LD. We also demonstrate that, once these variants are identified, target genes can be quickly identified using publicly available 3D chromatin data and additional functional assays such as CRISPRi.

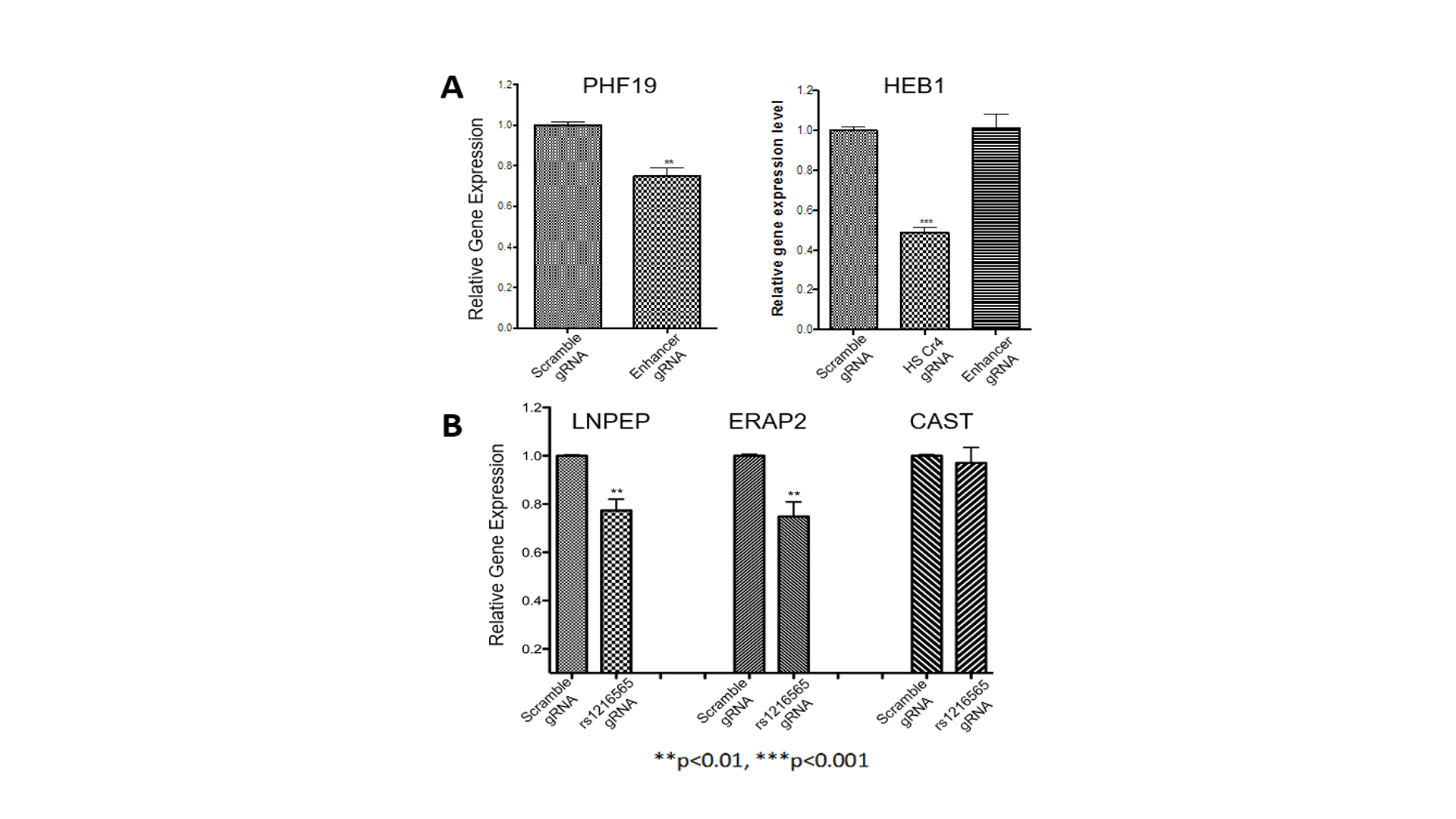
CRISPi identifies target genes of enhancers harboring JIA-associated variants detected on MPRA. (A). gRNAs used for epigenome editing experiments. The scrambled gRNA is a non-targeting negative control gRNA. For each experiment, 4 gRNAs were targeted to the functional regions of intergenic enhancers in the TRAF1 locus and the ERAP2/LNPEP locus. (B) Attenuation of the intergenic enhancer at the TRAF1 locus significantly reduced expression of PHF19, but not C5 (not shown). Off-target effects are monitored by showing that these gRNAs have no effect on HBE1 expression, but that gRNAs directed to an enhancer known to regulated HBE1 attenuates expression. Bar graphs summarize the results of 4 independent experiments. (C) Attenuating the intergenic enhancer in the LNPEP/ERAP2 locus reduces expression of both LNPEP and ERAP2, but not the adjacent gene, CAST, nor the HBE1 gene (not shown). In each experiment, scrambled versions of the gRNAs had no effect on expression of the putative targets. **p<0.01; ***p<0.001.
K. Jiang: None; t. liu: None; R. Tewhey: None; S. Kales: None; Y. Park: None; J. Jarvis: None.



