Poster Session A
Systemic lupus erythematosus (SLE)
Session: (0543–0581) SLE – Diagnosis, Manifestations, & Outcomes Poster I
0560: Genetic Determinants of Lupus Nephritis and Kidney Function in Systemic Lupus Erythematosus
Sunday, November 12, 2023
9:00 AM - 11:00 AM PT
Location: Poster Hall

Linda Hiraki, MD, FRCPC, ScD
The Hospital for Sick Children
Toronto, ON, CanadaDisclosure(s): No financial relationships with ineligible companies to disclose
Abstract Poster Presenter(s)
Magdalena Riedl Khursigara1, Nicholas Gold2, Thai-Son Tang3, Daniela Dominguez2, Marisa Klein-Gitelman4, Dafna Gladman5, Daniel Goldman6, Elizabeth Harvey2, Mariko Ishimori7, Caroline Jefferies7, Diane L. Kamen8, Sylvia Kamphuis9, Andrea Knight2, Chia-Chi Lee7, Deborah Levy2, Damien Noone2, Karen Onel10, Christine Peschken11, Michelle Petri6, Janet Pope12, Eleanor Pullenayegum2, Earl Silverman13, Zahi Touma1, Murray Urowitz14, Daniel Wallace7, Joan Wither15 and Linda Hiraki2, 1University of Toronto, Toronto, ON, Canada, 2The Hospital for Sick Children, Toronto, ON, Canada, 3University of Toronto, Mississauga, ON, Canada, 4Ann and Robert H. Lurie Children's Hospital of Chicago, Chicago, IL, 5Schroeder Arthritis Institute, Krembil Research Institute, Toronto Western Hospital, Department of Medicine, University of Toronto, Toronto, ON, Canada, 6Department of Medicine, Division of Rheumatology, Johns Hopkins University School of Medicine, Timonium, MD, 7Cedars-Sinai Medical Center, Los Angeles, CA, 8Medical University of South Carolina, Charleston, SC, 9Sophia Children’s Hospital, Rotterdam, Netherlands, 10Hospital for Special Surgery, New York, NY, 11University of Manitoba, Winnipeg, MB, Canada, 12University of Western Ontario, London, ON, Canada, 13Silverman, Toronto, ON, Canada, 14Schroeder Arthritis Institute, Krembil Research Institute; University of Toronto Lupus Clinic; Division of Rheumatology, Toronto, ON, Canada, 15University Health Network, Toronto, ON, Canada
Background/Purpose: Lupus nephritis (LN) is one of the most common and severe manifestation of SLE. Risk factors for lupus nephritis and renal function decline are not well understood. We completed a genome wide association study (GWAS) of LN in a multi-ethnic cohort of children and adults with SLE.
Methods: We included SLE patients from dedicated Lupus clinics and the SLICC cohort. All patients met ACR and/or SLICC SLE classification criteria and were genotyped on a multi-ethnic Illumina array. Ungenotyped SNPs were imputed to TopMed, and principal components for ancestral estimation were generated using both 1000 Genomes and GRAF-pop. LN was defined by SLE criteria and confirmed on kidney biopsy in 75%. Kidney function (estimated glomerular filtration rate, eGFR) was calculated using the Schwartz Bedside formula for measures < 18 years and CKD-EPI (without race) for >18 years of age and collected longitudinally over time. Wilcoxon rank sum or Chi-square tests were used to compare characteristics between LN and Non-LN patients. We completed GWAS of LN in marginal and multivariable adjusted regression models with principal components for ancestry, sex and cohort site.
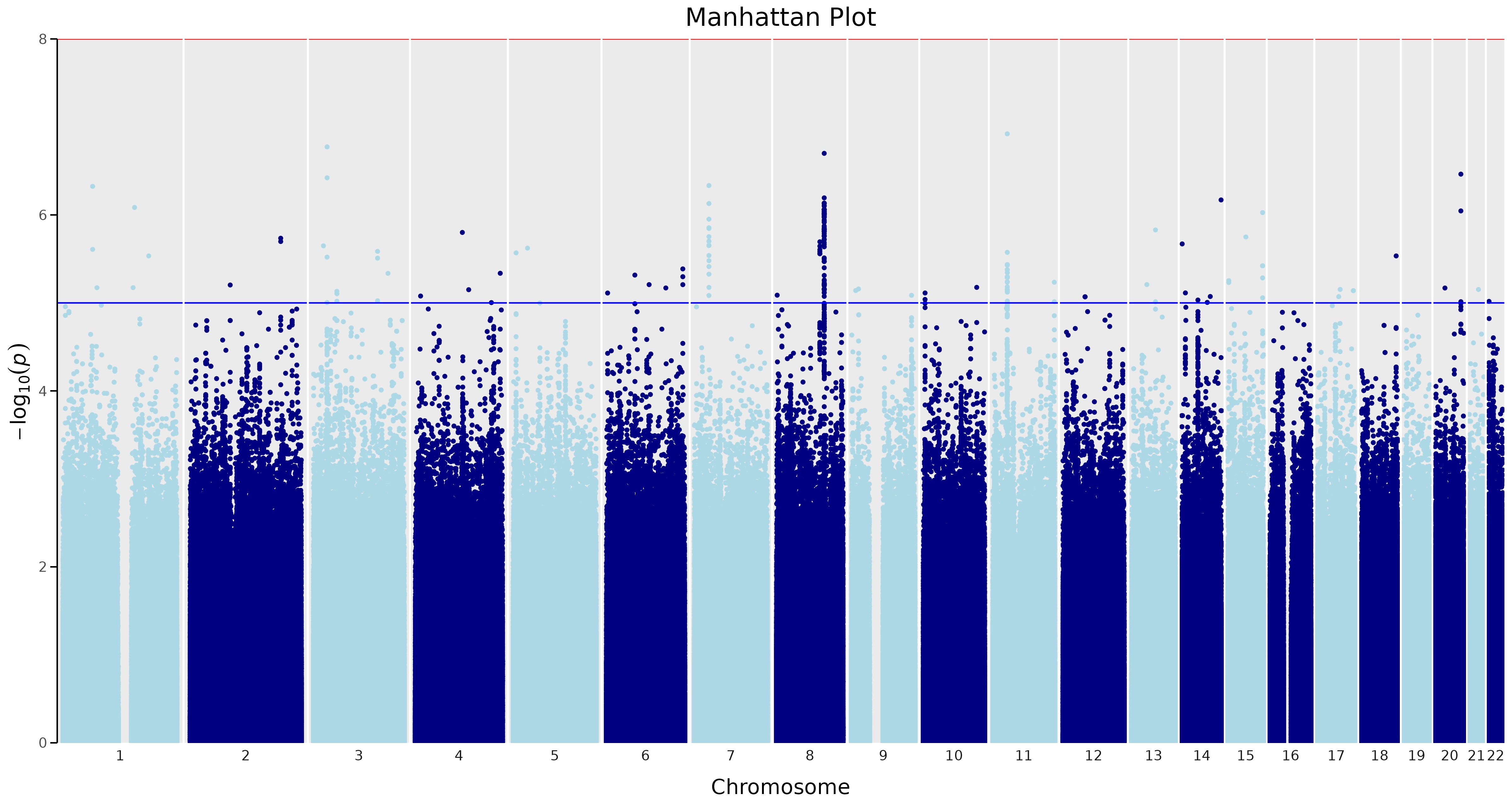
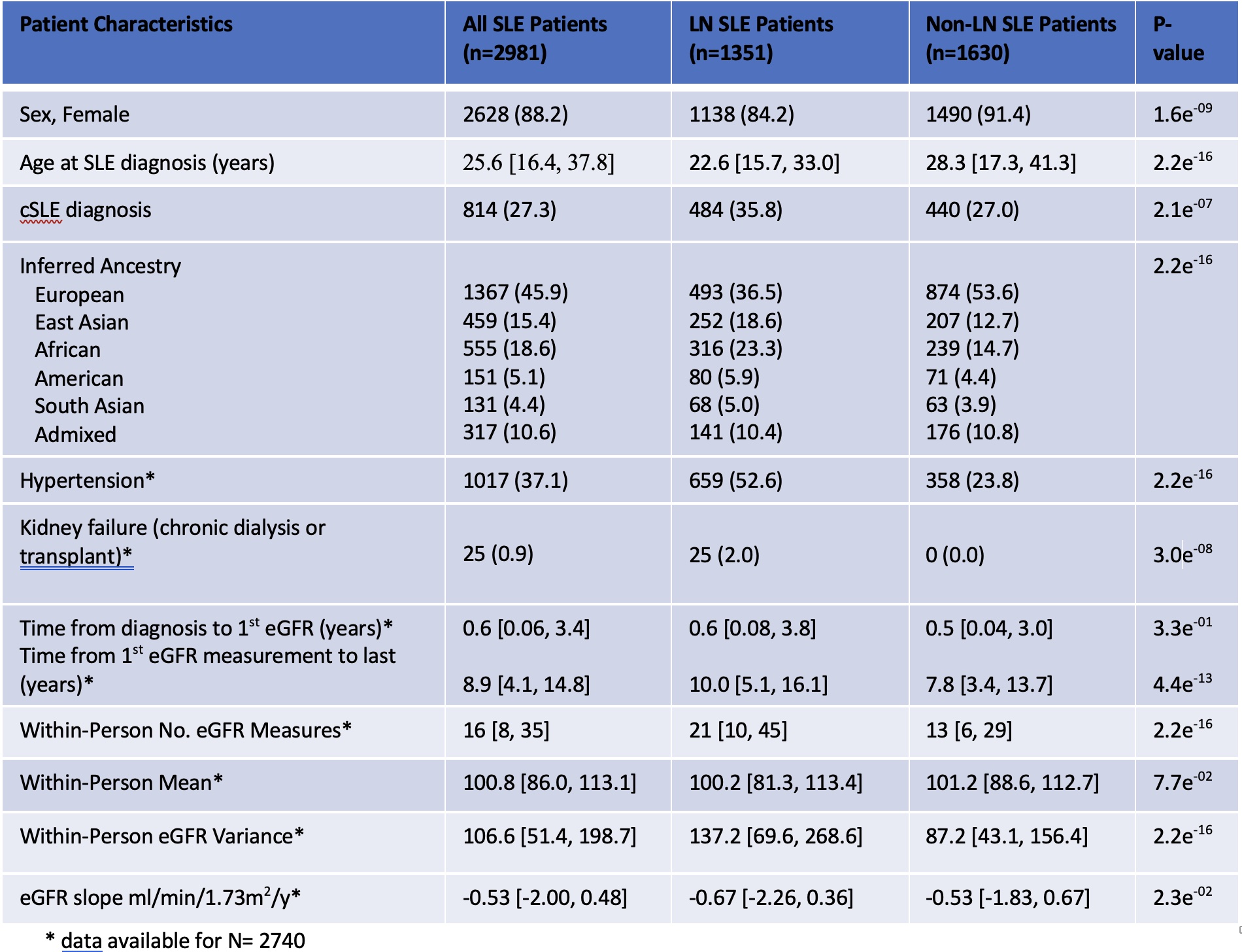
Results: The study included 2981 patients with SLE, 88% female, 46% of European ancestry and 27% childhood-onset SLE. LN was present in 45%. People with LN were younger at diagnosis, and more likely of African American or East Asian ancestry than those without LN. People with LN had significantly lower within-person mean eGFR, greater eGFR variability and slope over time compared to those without LN (Table). GWAS of LN demonstrated a peak on chromosome 8, intronic to ATP6V1C1, yet did not reach a genome-wide significance (p < 5x10-8).
Conclusion: A GWAS of LN in a multiethnic cohort of children and adults with SLE, did not identify a significant locus. The top signal was on chromosome 8, intronic to ATP6V1C1. We will complete GWAS of repeated eGFR measures, as it is a more informative outcome that we expect will improve power for detecting genetic loci for LN.
M. Riedl Khursigara: None; N. Gold: None; T. Tang: None; D. Dominguez: None; M. Klein-Gitelman: None; D. Gladman: AbbVie, 2, 5, Amgen, 2, 5, Bristol Myers Squibb, 2, Celgene, 2, 5, Eli Lilly, 2, 5, Galapagos, 2, Gilead Sciences, 2, Janssen, 2, 5, Novartis, 2, 5, Pfizer Inc, 2, 5, UCB, 2, 5; D. Goldman: None; E. Harvey: None; M. Ishimori: None; C. Jefferies: None; D. Kamen: None; S. Kamphuis: None; A. Knight: Pfizer, 6; C. Lee: None; D. Levy: None; D. Noone: None; K. Onel: None; C. Peschken: AstraZeneca, 2, 5, GSK, 2, 5, Roche, 1, 2; M. Petri: Alexion, 1, Amgen, 1, AnaptysBio, 1, Annexon Bio, 1, Argenx, 1, Arhros-Focus Med/Ed, 6, AstraZeneca, 1, 5, Aurinia, 1, 5, 6, Axdev, 1, Biogen, 1, Boxer Capital, 2, Cabaletto Bio, 2, Caribou Biosciences, 2, CVS Health, 1, Eli Lilly, 1, 5, Emergent Biosolutions, 1, Exagen, 5, Exo Therapeutics, 2, Gilead Biosciences, 2, GlaxoSmithKlein(GSK), 1, 5, 6, Horizon Therapeutics, 2, Idorsia Pharmaceuticals, 2, IQVIA, 1, Janssen, 1, 5, Kira Pharmaceuticals, 2, MedShr, 6, Merck/EMD Serono, 1, Momenta Pharmaceuticals, 2, Nexstone Immunology, 2, Nimbus Lakshmi, 2, Proviant, 2, Sanofi, 2, Sinomab Biosciences, 2, Thermofisher, 5, UCB, 2; J. Pope: AbbVie, 1, 2; E. Pullenayegum: None; E. Silverman: None; Z. Touma: AstraZeneca, 2, GSK, 2; M. Urowitz: None; D. Wallace: None; J. Wither: AstraZeneca, 1, 6, Pfizer, 12, Indirect salary support through a Chair award to the Division of Rheumatology at the University of Toronto; L. Hiraki: None.
Background/Purpose: Lupus nephritis (LN) is one of the most common and severe manifestation of SLE. Risk factors for lupus nephritis and renal function decline are not well understood. We completed a genome wide association study (GWAS) of LN in a multi-ethnic cohort of children and adults with SLE.
Methods: We included SLE patients from dedicated Lupus clinics and the SLICC cohort. All patients met ACR and/or SLICC SLE classification criteria and were genotyped on a multi-ethnic Illumina array. Ungenotyped SNPs were imputed to TopMed, and principal components for ancestral estimation were generated using both 1000 Genomes and GRAF-pop. LN was defined by SLE criteria and confirmed on kidney biopsy in 75%. Kidney function (estimated glomerular filtration rate, eGFR) was calculated using the Schwartz Bedside formula for measures < 18 years and CKD-EPI (without race) for >18 years of age and collected longitudinally over time. Wilcoxon rank sum or Chi-square tests were used to compare characteristics between LN and Non-LN patients. We completed GWAS of LN in marginal and multivariable adjusted regression models with principal components for ancestry, sex and cohort site.
Results: The study included 2981 patients with SLE, 88% female, 46% of European ancestry and 27% childhood-onset SLE. LN was present in 45%. People with LN were younger at diagnosis, and more likely of African American or East Asian ancestry than those without LN. People with LN had significantly lower within-person mean eGFR, greater eGFR variability and slope over time compared to those without LN (Table). GWAS of LN demonstrated a peak on chromosome 8, intronic to ATP6V1C1, yet did not reach a genome-wide significance (p < 5x10-8).
Conclusion: A GWAS of LN in a multiethnic cohort of children and adults with SLE, did not identify a significant locus. The top signal was on chromosome 8, intronic to ATP6V1C1. We will complete GWAS of repeated eGFR measures, as it is a more informative outcome that we expect will improve power for detecting genetic loci for LN.

Manhattan Plot for LN, adjusted for PC, sex and site

Table: Characteristics of LN and Non-LN patients
M. Riedl Khursigara: None; N. Gold: None; T. Tang: None; D. Dominguez: None; M. Klein-Gitelman: None; D. Gladman: AbbVie, 2, 5, Amgen, 2, 5, Bristol Myers Squibb, 2, Celgene, 2, 5, Eli Lilly, 2, 5, Galapagos, 2, Gilead Sciences, 2, Janssen, 2, 5, Novartis, 2, 5, Pfizer Inc, 2, 5, UCB, 2, 5; D. Goldman: None; E. Harvey: None; M. Ishimori: None; C. Jefferies: None; D. Kamen: None; S. Kamphuis: None; A. Knight: Pfizer, 6; C. Lee: None; D. Levy: None; D. Noone: None; K. Onel: None; C. Peschken: AstraZeneca, 2, 5, GSK, 2, 5, Roche, 1, 2; M. Petri: Alexion, 1, Amgen, 1, AnaptysBio, 1, Annexon Bio, 1, Argenx, 1, Arhros-Focus Med/Ed, 6, AstraZeneca, 1, 5, Aurinia, 1, 5, 6, Axdev, 1, Biogen, 1, Boxer Capital, 2, Cabaletto Bio, 2, Caribou Biosciences, 2, CVS Health, 1, Eli Lilly, 1, 5, Emergent Biosolutions, 1, Exagen, 5, Exo Therapeutics, 2, Gilead Biosciences, 2, GlaxoSmithKlein(GSK), 1, 5, 6, Horizon Therapeutics, 2, Idorsia Pharmaceuticals, 2, IQVIA, 1, Janssen, 1, 5, Kira Pharmaceuticals, 2, MedShr, 6, Merck/EMD Serono, 1, Momenta Pharmaceuticals, 2, Nexstone Immunology, 2, Nimbus Lakshmi, 2, Proviant, 2, Sanofi, 2, Sinomab Biosciences, 2, Thermofisher, 5, UCB, 2; J. Pope: AbbVie, 1, 2; E. Pullenayegum: None; E. Silverman: None; Z. Touma: AstraZeneca, 2, GSK, 2; M. Urowitz: None; D. Wallace: None; J. Wither: AstraZeneca, 1, 6, Pfizer, 12, Indirect salary support through a Chair award to the Division of Rheumatology at the University of Toronto; L. Hiraki: None.



