Abstract Session
Osteoarthritis (OA) and related disorders
Session: Abstracts: Osteoarthritis II: Novel Insights from Observational Studies (2467–2472)
2468: Association Between Gut Microbiome-related Metabolites and Symptomatic Hand Osteoarthritis in Two Independent Cohorts
Tuesday, November 14, 2023
2:15 PM - 2:25 PM PT
Location: Room 6F/6C
- JW
Jie Wei, PhD (she/her/hers)
Health Management Center, Xiangya Hospital Central South University
Changsha, ChinaDisclosure information not submitted.
Presenting Author(s)
Jie Wei1, Zidan Yang2, Jiatian Li3, Yuqing Zhang4, Weiya Zhang5, Michael Doherty5, Tuo Yang6, Yuanheng Yang3, Hui Li3, Yilun Wang3, Ziying Wu3, Changjun Li7, Guanghua Lei3 and Chao Zeng3, 1Health Management Center, Xiangya Hospital Central South University, Changsha, China, 2Key Laboratory of Aging-related Bone and Joint Diseases Prevention and Treatment, Ministry of Education, Xiangya Hospital, Central South University, Changsha, China, 3Department of Orthopaedics, Xiangya Hospital, Central South University, Changsha, China, 4Division of Rheumatology, Allergy, and Immunology, Department of Medicine, Massachusetts General Hospital, Boston, MA, 5Academic Rheumatology, Clinical Sciences Building, University of Nottingham, City Hospital, Nottingham, United Kingdom, 6Department of Health Management Center, Xiangya Hospital, Central South University, Changsha, China, 7National Clinical Research Center for Geriatric Disorders, Xiangya Hospital, Central South University, Changsha, China
Background/Purpose: Since gut microbiome dysbiosis can cause inflammatory disorders by affecting host metabolism, we postulate that the gut microbiome and related metabolites could play a role in the pathogenesis of hand osteoarthritis. We characterised gut microbiome-related metabolites in people with symptomatic hand OA (SHOA) in two independent cohorts.
Methods: Using data collected from a large-sample community-based observational study (discovery cohort) (n=1,359), the Xiangya OA (XO) Study, we first performed shotgun metagenomic sequencing of stool samples for gut microbial function. We then performed targeted metabolomics analysis of plasma samples for metabolites related to altered microbial function in SHOA and compared the metabolites between participants with and those without SHOA. We also applied a multi-omics analysis approach to examine the associations between SHOA-related microbial function and SHOA-related plasma metabolites. Finally, we then replicated the findings from the XO Study among 71 SHOA patients and 71 age (±1 year), sex and BMI (±2 kg/m2) matched controls (validation cohort) (Figure 1).
Results: In the discovery cohort, compared with those without SHOA (n=70), participants with SHOA had significantly altered microbial functions related to tryptophan metabolism. Then we measured the plasma tryptophan metabolites and found that participants with SHOA had higher levels of 5-hydroxytryptophol and 5-hydroxyindoleacetic acid, but lower levels of indole-3-lactic acid (ILA), 3-hydroxyanthranilic acid and skatole (Figure 2). The multivariable-adjusted odd ratios of SHOA per one log10 unit increase for 5-HIAA, 5-HTOL, ILA, skatole, and 3-HAA were 8.97 (95% CI: 2.37 to 33.98), 3.44 (95% CI: 1.44 to 8.18), 0.19 (95% CI: 0.04 to 0.99), 0.51 (95% CI: 0.28 to 0.90) and 0.36 (95% CI: 0.19 to 0.70), respectively. Moreover, the multi-omics analysis suggested that the lower level of ILA in SHOA was related to the microbial function of tryptophan biosynthesis, which was further correlated with the microbiome species Bacteroides A mediterraneensis (Figure 3). Finally, we verified the associations between plasma tryptophan metabolites and SHOA in the validation cohort. Consistent with the findings in the discovery cohort, we observed a lower level of the ILA in the SHOA group compared with controls, with the multivariable adjusted ORs of SHOA per one log10 unit increase for ILA being 0.03 (95% CI: 0.002 to 0.46).
Conclusion: Alterations of the microbial function of tryptophan biosynthesis and tryptophan metabolites, especially lower levels of ILA, are associated with SHOA. Our novel findings suggest potential pathways and targets for further treatment of SHOA.
.jpg)

J. Wei: None; Z. Yang: None; J. Li: None; Y. Zhang: None; W. Zhang: None; M. Doherty: None; T. Yang: None; Y. Yang: None; H. Li: None; Y. Wang: None; Z. Wu: None; C. Li: None; G. Lei: None; C. Zeng: None.
Background/Purpose: Since gut microbiome dysbiosis can cause inflammatory disorders by affecting host metabolism, we postulate that the gut microbiome and related metabolites could play a role in the pathogenesis of hand osteoarthritis. We characterised gut microbiome-related metabolites in people with symptomatic hand OA (SHOA) in two independent cohorts.
Methods: Using data collected from a large-sample community-based observational study (discovery cohort) (n=1,359), the Xiangya OA (XO) Study, we first performed shotgun metagenomic sequencing of stool samples for gut microbial function. We then performed targeted metabolomics analysis of plasma samples for metabolites related to altered microbial function in SHOA and compared the metabolites between participants with and those without SHOA. We also applied a multi-omics analysis approach to examine the associations between SHOA-related microbial function and SHOA-related plasma metabolites. Finally, we then replicated the findings from the XO Study among 71 SHOA patients and 71 age (±1 year), sex and BMI (±2 kg/m2) matched controls (validation cohort) (Figure 1).
Results: In the discovery cohort, compared with those without SHOA (n=70), participants with SHOA had significantly altered microbial functions related to tryptophan metabolism. Then we measured the plasma tryptophan metabolites and found that participants with SHOA had higher levels of 5-hydroxytryptophol and 5-hydroxyindoleacetic acid, but lower levels of indole-3-lactic acid (ILA), 3-hydroxyanthranilic acid and skatole (Figure 2). The multivariable-adjusted odd ratios of SHOA per one log10 unit increase for 5-HIAA, 5-HTOL, ILA, skatole, and 3-HAA were 8.97 (95% CI: 2.37 to 33.98), 3.44 (95% CI: 1.44 to 8.18), 0.19 (95% CI: 0.04 to 0.99), 0.51 (95% CI: 0.28 to 0.90) and 0.36 (95% CI: 0.19 to 0.70), respectively. Moreover, the multi-omics analysis suggested that the lower level of ILA in SHOA was related to the microbial function of tryptophan biosynthesis, which was further correlated with the microbiome species Bacteroides A mediterraneensis (Figure 3). Finally, we verified the associations between plasma tryptophan metabolites and SHOA in the validation cohort. Consistent with the findings in the discovery cohort, we observed a lower level of the ILA in the SHOA group compared with controls, with the multivariable adjusted ORs of SHOA per one log10 unit increase for ILA being 0.03 (95% CI: 0.002 to 0.46).
Conclusion: Alterations of the microbial function of tryptophan biosynthesis and tryptophan metabolites, especially lower levels of ILA, are associated with SHOA. Our novel findings suggest potential pathways and targets for further treatment of SHOA.
.jpg)
Figure 1. Summary of the present study. SHOA, symptomatic hand osteoarthritis; KEGG, Kyoto Encyclopedia of Genes and Genomes.

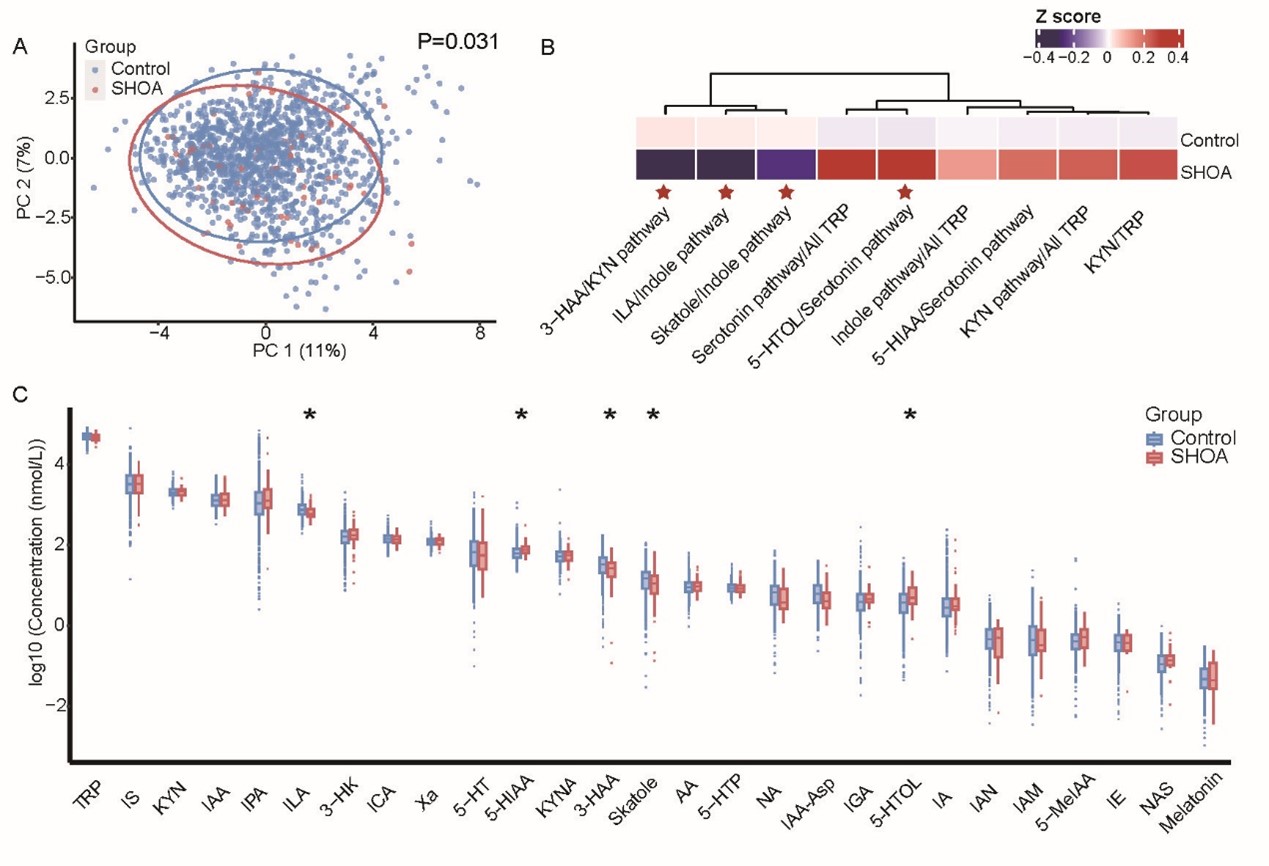
Figure 2. Associations of plasma tryptophan metabolites with SHOA in the discovery cohort. (A) Principal Component Analysis (PCA) plot compared the composition of plasma tryptophan metabolites between participants with SHOA and those without SHOA. Associations of ratios of tryptophan metabolites concentration in their related pathway (B) and individual plasma tryptophan metabolites (C) with SHOA adjusting for age, sex, body mass index, smoking status, alcohol consumption, and frequency of dietary intake of meat/eggs, dairy, and vegetables. Significant ratios are marked with red stars. SHOA, symptomatic hand osteoarthritis; TRP, tryptophan; KYN, kynurenine; ILA, indole-3-lactic acid; 3-HAA, 3-hydroxyanthranilic acid; 5-HIAA, 5-hydroxyindoleacetic acid; 5-HTOL, 5-hydroxytryptophol; 3-HK; 3-hydroxykynurenine; 5-HT, serotonin; 5-HTP, L-5-hydroxytryptophan; 5-MeIAA; 5-methoxy-3-indoleacetic acid; AA, anthranilic acid; IA; indole acrylic acid; IAA, indole-3-acetic acid; IAA-Asp, indole-3-acetyl-aspartate; IAM, indole-3-acetamide; IAN, indole-3-acetonitrile; ICA, indole-3-carboxaldehyde; IE, indole ethanol/tryptophol; IGA, 3-indoleglyoxylic acid; IPA, 3-indolepropionic acid; IS, indoxylsulfate; KYNA, kynurenic acid; NA, nicotinic acid; NAS, N-acetyl-5-hydroxytryptamine; Xa, xanthurenic acid.

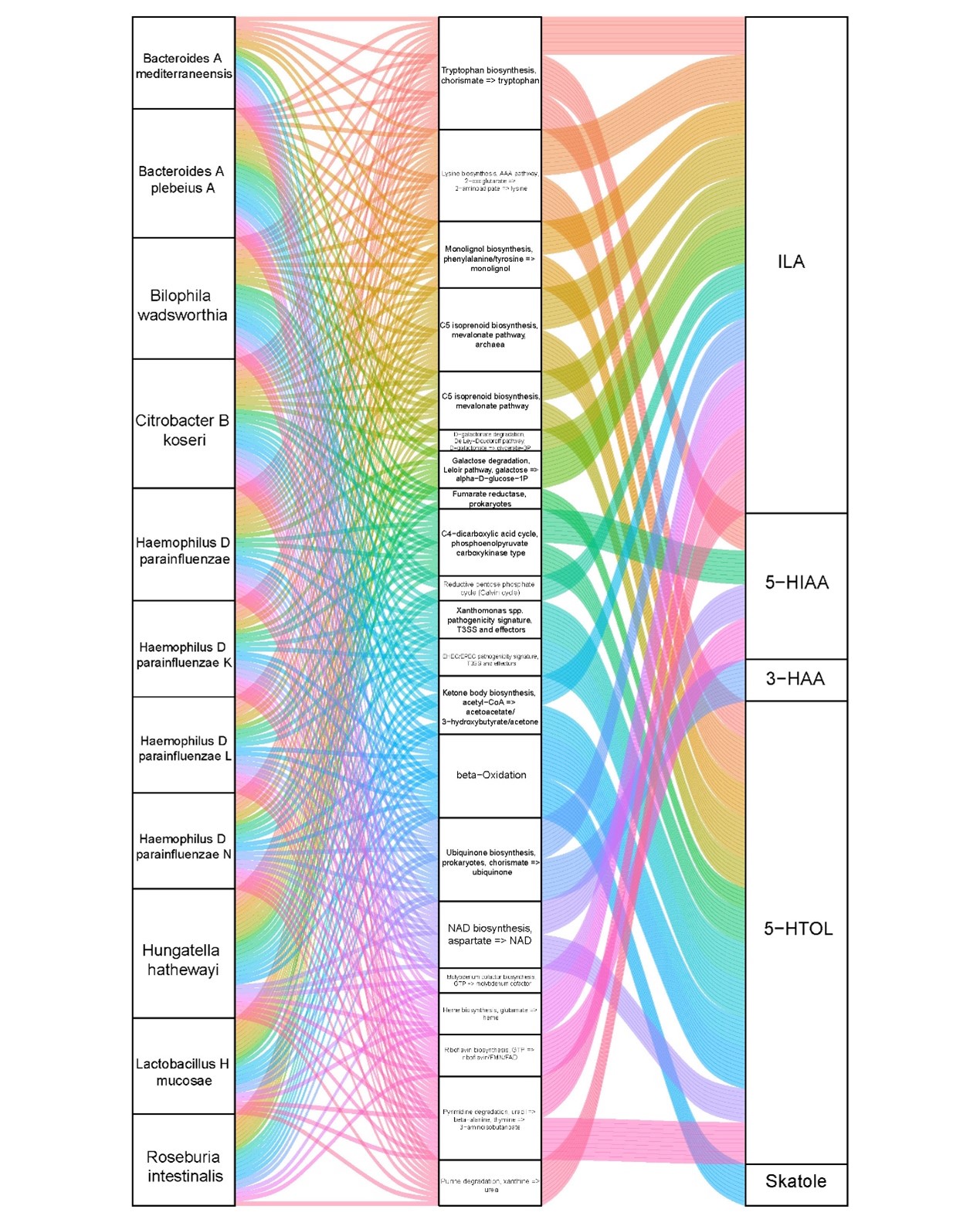
Figure 3. Correlations of microbial species to the microbial modules, and correlations of microbial modules to the tryptophan metabolites. The Sankey plot shows the significant correlations of classified microbial species with KEGG modules and KEGG modules with differential tryptophan metabolites. Shown are microbial species (left), KEGG modules (middle), and plasma tryptophan metabolites (right). The curved lines connecting the panels indicate the correlations, with colors corresponding to different modules. ILA, indole-3-lactic acid; 5-HIAA, 5-hydroxyindoleacetic acid; 3-HAA, 3-hydroxyanthranilic acid; 5-HTOL, 5-hydroxytryptophol.
J. Wei: None; Z. Yang: None; J. Li: None; Y. Zhang: None; W. Zhang: None; M. Doherty: None; T. Yang: None; Y. Yang: None; H. Li: None; Y. Wang: None; Z. Wu: None; C. Li: None; G. Lei: None; C. Zeng: None.



